Advanced information
![]()
Advanced information:
RNA INTERFERENCE [pdf]
RNA INTERFERENCE
This year’s Nobel Prize in Physiology or Medicine is shared by Professor Andrew Z. Fire at Stanford University, California, USA, and Professor Craig C. Mello at the University of Massachusetts Medical School in Worcester, USA. They receive the prize for their discovery that double-stranded RNA triggers suppression of gene activity in a homology-dependent manner, a process named RNA interference (RNAi). Their discovery revealed a new mechanism for gene regulation, and the biochemical machinery involved plays a key role in many essential cellular processes. Double-stranded RNA synthesized within the cell can reduce or abolish gene activity by RNAi-like mechanisms. This control system for gene expression has proven to be important for both the development of an organism and the physiological functions of cells and tissues. Furthermore, RNAi protects against RNA virus infections, especially in plants and invertebrate animals, and secures genome stability by keeping mobile elements silent. Today, double-stranded RNA is used as a powerful tool to experimentally elucidate the function of essentially any gene in a cell. The discovery of RNAi has already had an immense impact on biomedical research and will most likely lead to novel medical applications in the future.
Introduction
The gene expression process is of fundamental importance for all living organisms. Most genes reside in the chromosomes located in the cell nucleus and express themselves via proteins synthesised in the cytoplasm. The genetic material was identified as deoxyribonucleic acid (DNA) in 1944 (ref. 1) and the double-helical nature of DNA was revealed in 1953 (by Francis Crick, James Watson and Maurice Wilkins; Nobel Prize in Physiology or Medicine in 1962). At that time, the main problem outstanding was how DNA in the cell nucleus could govern protein synthesis in the cytoplasm. It was proposed that another nucleic acid, single-stranded ribonucleic acid (RNA), acts as an intermediary in the process, and the so-called Central Dogma was formulated, i.e. the idea that the genetic information is transcribed from DNA to RNA and then translated from RNA into protein. The RNA carrying the genetic information was first believed to be the RNA in ribosomes; for several years the hypothesis was formulated as “one gene-one ribosome-one protein”. In 1961, Francois Jacob and Jacques Monod presented a visionary gene control model, for which they received the Nobel Prize in Physiology or Medicine in 1965 together with André Lwoff. In their model, they proposed that the gene is transcribed into a specific RNA species, messenger RNA (mRNA). Soon afterwards it was demonstrated that a short-lived, non-ribosomal RNA directs the synthesis of proteins. Subsequently, Marshall Nirenberg and Gobind Khorana broke the genetic code and could assign code words (codons; triplets of nucleotides) to the twenty amino acids (they received the Nobel Prize in Physiology or Medicine in 1968 together with Robert Holley). Francis Crick predicted that an RNA molecule could act as an adaptor between mRNA and the amino acid, and a short, stable RNA, transfer RNA (tRNA) was soon identified as the predicted adaptor.
For many years, messenger RNA was believed to correspond to an uninterrupted nucleotide sequence in the DNA. It therefore came as a complete surprise when Phillip Sharp and Richard Roberts showed in 1977 that the mRNA sequence could be distributed discontinuously in the genome (the split gene concept; Nobel Prize in 1993). It was known that long RNA molecules (pre-mRNA, heterogeneous nuclear RNA) are trimmed to much shorter mature mRNAs, and Sharp and Roberts therefore suggested that the mRNA sequences, the exons, are likely to be cut out from the primary transcript and spliced, while the intervening sequences, the introns, are degraded. It was immediately realised that the discontinuous arrangement of the mRNA sequences in DNA had important evolutionary implications. In addition, the RNA splicing process could generate different mRNAs, and more than one protein can emanate from the primary transcript (alternative splicing).
The discovery that RNA can act as a catalyst gave a radically new perspective on the roles of RNA (Nobel Prize in Chemistry to Sidney Altman and Thomas Cech in 1989). It was soon revealed that RNA is able to catalyse its own replication and the synthesis of other RNA molecules (the ribozyme concept), which led to the idea that RNA was the first genetic material on earth. An “RNA world” is believed to have existed before DNA took over the role of being the key genetic material, and RNA was relegated to the role of messenger between DNA and protein. Not only did the discovery of catalytic RNA have evolutionary implications, but it also suggested that RNA could play a more active role in gene expression than earlier realised. It is now well established that ribosomal RNA catalyses peptide bond formation during translation.
A large number of small RNA molecules work in conjunction with proteins in ribonucleoprotein (RNP) complexes. There are non-coding RNA molecules that affect transcription (e.g. human 7SK snRNA bound to elongation factors), translation (e.g. SRP RNA in the signal recognition particle), replication (e.g. telomerase RNA) and chromosome structure (e.g. XIST RNA causing X chromosome inactivation). Others regulate RNA processing (e.g. M1 RNA in RNase P, snRNAs and snoRNAs) and RNA editing (guide RNAs). These various RNP particles are now being extensively investigated to understand their specific roles in the cell.
In the early 1980s it was revealed in Escherichia coli that small RNA molecules (about 100 nucleotides in length) can bind to a complementary sequence in mRNA and inhibit translation2,3. Today, about 25 cases of regulatory trans-acting antisense RNAs are known in E. coli4. Regulation of translation by antisense RNA also occurs in eukaryotes as was first demonstrated in 1993 when genes governing the development of the worm Caenorhabditis elegans were studied5,6. For many years, this thoroughly documented case of posttranscriptional regulation was regarded as an oddity. The mechanism received more attention when a second example of a small regulatory RNA was found in C. elegans7, because in this case similar sequences were also present in other species. However, the situation changed drastically when a large number of small RNA molecules, called microRNAs (miRNAs), were revealed in 2001 (ref. 8-10).
Prior to the discovery of RNA interference, a phenomenon called gene (or RNA) silencing was described in plants. It was noted in experiments around 1990 that a cloned gene incorporated into the genome (a transgene) could not only induce or stimulate gene activity but could also inhibit the expression of homologous sequences, a phenomenon called homology-dependent gene silencing. The inhibition of gene activity could take place at the transcriptional level (transcriptional gene silencing, TGS)11-13, or at the posttranscriptional level (posttranscriptional gene silencing, PTGS)14-18. A PTGS-like process called quelling was also established in the fungus Neurospora crassa19. Analyses of viral infection in plants gave further insight into the mechanism of PTGS20-21. However, although it was evident that RNA played a key role in gene silencing, the phenomenon remained enigmatic until the discovery of RNA interference provided a most unexpected explanation with many profound consequences.
The discovery of RNA interference
Andrew Fire and Craig Mello published their break-through study on the mechanism of RNA interference in Nature in 1998 (ref. 22). It was earlier known that antisense RNA23, but remarkably also sense RNA24 could silence genes, but the results were inconsistent and the effects usually modest. However, due to the fact that both sense and antisense RNA could cause silencing Mello argued that the mechanism could not just be a pairing of antisense RNA to mRNA, and he coined the term RNA interference for the unknown mechanism25.
In their Nature paper, Fire and Mello tested the phenotypic effect of RNA injected into the worm C. elegans. They established that annealed sense/antisense RNA, but neither antisense nor sense RNA alone, caused the predicted phenotype (Fig. 1). Furthermore, only injection of double-stranded RNA (dsRNA) led to an efficient loss of the target mRNA (Fig. 2). Fire and Mello could present a series of straightforward conclusions in their study. The main results can be summed up as follows: First, silencing was triggered efficiently by injected dsRNA, but weakly or not at all by sense or antisense single-stranded RNAs. Second, silencing was specific for an mRNA homologous to the dsRNA; other mRNAs were unaffected. Third, the dsRNA had to correspond to the mature mRNA sequence; neither intron nor promoter sequences triggered a response. This indicated a posttranscriptional, presumably cytoplasmic mechanism. Fourth, the targeted mRNA disappeared suggesting that it was degraded. Fifth, only a few dsRNA molecules per cell were sufficient to accomplish full silencing. This indicated that the dsRNA was amplified and/or acted catalytically rather than stoichiometrically. Sixth, the dsRNA effect could spread between tissues and even to the progeny, suggesting a transmission of the effect between cells. Furthermore, Fire and Mello made the remark that RNAi could provide an explanation for a phenomenon studied in plants for several years: posttranscriptional gene silencing (PTGS). Finally, they ended their paper by speculating about the possibility that “dsRNA could be used by the organism for physiological gene silencing”.
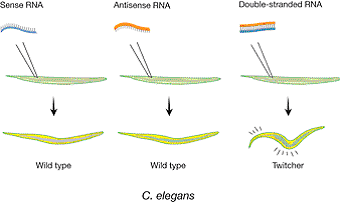 |
| Figure 1. Phenotypic effect after injection of single-stranded or double-stranded unc-22 RNA into the gonad of C. elegans. The unc-22 gene encodes a myofilament protein. Decrease in unc-22 activity is known to produce severe twitching movements. Injected double-stranded RNA, but not single-stranded RNA, induced the twitching phenotype in the progeny. |
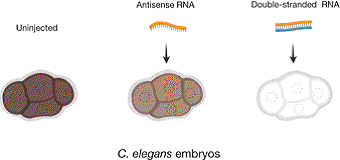
Figure 2. The effect on mex-3 mRNA content in embryos after injection of single-stranded or double-stranded mex-3 RNA into the gonad of C. elegans. mex-3 mRNA is abundant in the gonads and early embryos. The mRNA was lost after injection of double-stranded RNA, while injection of antisense RNA only reduced the content of mRNA to some extent. The extent of brown colour reflects the amount of mRNA present.
In their Nature paper, Fire and Mello did not take a firm stand on the issue of whether dsRNA acts via a transcriptional or posttranscriptional mechanism. However, in a follow-up study published in PNAS the same year, Fire provided good evidence for the view that mRNA is the target for dsRNA (recognition via complementary strands), and that the targeted mRNA is degraded prior to translation, i.e. dsRNA exerts its effect at the posttranscriptional level26. He also presented a specific model showing how double-stranded RNA could function in a catalytic manner to target homologous mRNAs for degradation. This model differed strikingly from the simple antisense model of that time, which only predicted interaction between an interfering single-stranded RNA molecule and mRNA. It can be added that in this PNAS paper, Fire also indicated the possibility that the RNAi mechanism could be a specific “tactical” approach to viral defence in lower organisms (to be compared with the global interferon response in mammals).
Within a year, the presence of RNAi had been documented in many other organisms, including fruit flies, trypanosomes, plants, planaria, hydra and zebrafish27. In initial experiments with mammalian cultured cells, it was not possible to elicit a potent and specific RNAi response because of a predominant nonspecific physiological reaction of these cells to long dsRNA. However, when the cells were exposed to a short, 21 nucleotide long, dsRNA, an efficient targeted silencing was also obtained in these cells28. Thus, the generality of the RNAi phenomenon among eukaryotes was proven very rapidly; a remarkable exception is the budding yeast, Saccharomyces cerevisiae.
The revelation of the RNAi mechanism
Shortly after the discovery of RNAi, it was shown that PTGS in plants is correlated to the presence of a population of small RNAs (about 25 nucleotides long), and that this RNA contains both sense and antisense RNA sequences29. It was proposed that this RNA is the determinant of PTGS. Earlier, but subsequent to the discovery of RNAi in animal cells, it had been shown that RNA in double-stranded form also initiates PTGS in plants30.
The biochemistry of RNAi was further elucidated in an in vitro system based on Drosophila embryo extracts27. It could be shown that dsRNA is processed to 21-23 nucleotide long dsRNA fragments31, which was in good agreement with the data for PTGS in plants29. It was proposed that this short dsRNA, siRNA (small interfering RNA), guided the cleavage of mRNA. Subsequently, Fire and Mello were able to follow the process in vivo32. They established that long dsRNA is cleaved to small RNA (about 25 nucleotides long), and that antisense RNA triggers degradation of mRNA via base-pairing to mRNA. Thus, the trigger, dsRNA, had then been connected to the low molecular weight effector.
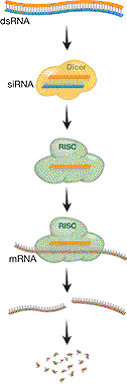 |
| Figure 3. The RNA interference process and the biochemical machinery involved. Double-stranded RNA is cut into short pieces (siRNA) by the endonuclease Dicer. The antisense strand is loaded into the RISC complex and links the complex to the mRNA strand by base-pairing. The RISC complex cuts the mRNA strand, and the mRNA is subsequently degraded. |
The molecular machinery involved in RNAi was subsequently revealed (Fig. 3). In an in vitro system, built on Drosophila cultured cells, it was demonstrated that a large complex, called RISC (RNA-induced silencing complex) is targeted to the mRNA via a short antisense RNA, and that mRNA is cleaved and subsequently degraded33. Subsequently, it has been shown that RISC contains at least one member of the argonaute protein family, which is likely to act as an endonuclease and cut the mRNA (nowadays often referred to as the Slicer function). It was also demonstrated that a ribonuclease III-like nuclease, called Dicer, is responsible for the processing of dsRNA to short RNA34. In certain systems, in particular plants, worms and fungi, an RNA dependent RNA polymerase (RdRP) plays an important role in generating and/or amplifying siRNA35.
Thus, in just a few years a vast amount of information accumulated on the specific proteins and protein complexes involved in RNAi and on the molecular details of the particular steps in the process36-39.
Significance of the discovery of RNAi
It was evident from the very beginning that the significance of the discovery of RNAi would be exceptional. The far-reaching consequences of the discovery can be summed up as follows (Fig. 4):
1. RNAi protects against viral infections: The finding of Fire and Mello that cells can process injected dsRNA and eliminate homologous single-stranded RNA suggested that RNAi could constitute a defence mechanism against viral attacks. It had earlier been shown that plant cells have an efficient defence against viruses based on the PTGS phenomenon40,41. When it became apparent that PTGS is the plant equivalent to RNAi, this early work in plants supported the proposition that RNAi is involved in protecting cells from viral attacks. Today, we know that this anti-viral mechanism is at work in plants, worms and flies, whereas it is still unclear how relevant it is for vertebrates, including man.
2. RNAi secures genome stability by keeping mobile elements silent: It was proposed early on that RNAi/PTGS in C. elegans and plants could block the action of transposons (mobile elements in the genome). Subsequently, it could be shown that when components of the RNAi machinery are mutated in C. elegans, transposons are activated and the mobile elements cause disturbances in the function of the genome42,43. It has been proposed that in transposon-containing regions of the genome both DNA strands are transcribed, dsRNA is formed, and the RNAi process eliminates these undesirable products. As short dsRNAs can also operate directly on chromatin and suppress transcription (see below), this would be another mode to keep transposons inactive (see below under 4). Even if the mechanisms are not yet fully revealed, it is clear that if the RNAi machinery is not efficient, the transposons are not kept under control and can start to jump and cause deleterious effects in the genome.
It has been argued that RNA silencing could represent an “immune defence” of the genome44. Close to 50% of our genome consists of viral and transposon elements that have invaded the genome in the course of evolution. The RNAi machinery can recognize invading double-stranded viral RNA (or the double-stranded replicative form of the viral RNA) and suppress the infection by degradation of the RNA. The RNAi system thus shares important features with the vertebrate immune system: it recognizes the invading parasite (dsRNA), raises an initial response and subsequently amplifies the response to eliminate the foreign element.
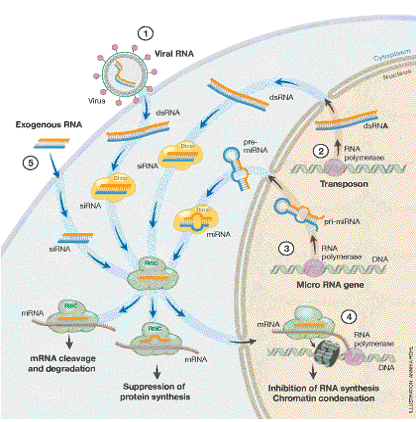
Figure 4. Cellular processes dependent on the RNAi machinery. The Dicer and RISC complexes play a central role in the destruction of invading viral RNA (1), the elimination of transcripts from mobile elements (transposons) and repetitive DNA (2), the block of protein synthesis brought about by small RNAs generated within the cell (3), and the RNAi-mediated suppression of transcription (4). The machinery is also utilized when siRNA is introduced into the cell experimentally to inhibit the activity of specific genes (5). The figure is schematic, and the Dicer and RISC complexes can vary dependent on cellular process.
3. RNAi-like mechanisms repress protein synthesis and regulate the development of organisms: Soon after the discovery that short RNA is the effector of RNAi, it was shown that there is a class of endogenous RNA molecules of the same size in worms, flies, mice and humans; this small RNA was called microRNA (miRNA)8-10. Plants also contain this class of endogenous RNA45. The revelation of miRNA led to intense research on the nature of these RNA molecules. The C. elegans lin-4 (ref. 5) and let-7 (ref. 7) RNAs were regarded as prototypes, and examples of similar cases were soon revealed in several organisms46. The small miRNAs are processed from larger hairpin-like precursors by an RNAi-like machinery47,48 (Fig. 4). The miRNAs can regulate gene expression by base-pairing to mRNA, which results in either degradation of the mRNA or suppression of translation. Today, it is estimated that there are about 500 miRNAs in mammalian cells, and that about 30% of all genes are regulated by miRNAs. It is known that miRNAs play an important role during development in plants, C. elegans and mammals. Thus, the miRNA-dependent control of gene expression represents a new major principle of gene regulation. However, the full significance of small regulatory RNAs is probably still not apparent.
4. RNAi-like mechanisms keep chromatin condensed and suppress transcription: It was known from work in plants that gene silencing could take place at the transcriptional level (TGS). After the discovery of RNAi, it was soon shown that TGS in plants operates via RNAi-like mechanisms49,50. In the fission yeast Schizosaccharomyces pombe51,52, and later on in Drosophila and vertebrates, it was found that similar processes keep heterochromatic regions condensed and transcriptionally suppressed. In addition, the RNAi-like machinery regulates the activity of genes in the immediate vicinity of the condensed blocks of chromatin. The phenomenon is still not understood at the molecular level although histone modifications, binding of specific chromatin condensing proteins (HP1), and DNA methylation all play important roles46. It is, however, evident that this action on chromatin is most important for proper functioning of the genome and for maintenance of genome integrity.
5. RNAi offers a new experimental tool to repress genes specifically. The targeted action of RNAi immediately suggested that this phenomenon could be utilized as a general method to suppress specific genes and look for the resulting phenotypic effect. It was also soon evident that this could be accomplished in such an efficient manner that essentially any gene in an organism could be studied functionally. After the initial work in C. elegans, this technical approach could be applied to cells from almost all organisms, including mammalian cells. This targeted gene silencing by RNAi has already had a tremendous impact in studies of the function of individual genes. It is now exploited not only in cultured cells but also in transgenic organisms. DNA constructs are introduced into the organisms under appropriate promoter control, and dsRNA hairpin structures are produced and further processed to achieve specific effects on gene regulation.
6. RNAi might be a useful approach in future gene therapy. The possibility to achieve RNAi-governed gene regulation in transgenic organisms has stimulated many explorations of whether this would be a useful option for medical therapy53,54. Promising results have been reported in several animal models55-58 and even in recent clinical trials, but it is too early to predict the outcome of these challenging efforts.
Conclusions
The discovery that cells have a special mechanism for suppressing the expression of homologous genes by recognizing and processing double-stranded RNA was totally unexpected and has dramatically expanded our knowledge of gene control. Remarkably, the RNAi machinery can handle double-stranded RNA entering the cell as well as double-stranded RNA generated within the cell. The development of an organism and proper function of its cells and tissues are dependent on an intact RNAi machinery. Infection by RNA viruses can be blocked by RNAi, especially in plants and lower animals, and foreign elements in the genome (viruses and transposons) can be kept silent. Finally, the discovery of RNAi has not only provided us with a powerful new experimental tool to study the function of genes but also raises expectations about future applications of RNAi in medicine.
References
1. Avery, OT, MacLeod, CM and McCarty, M (1944). Studies on the chemical nature of the substance inducing transformation of Pneumococcal types. J. Exp. Med. 79, 137-158.
2. Mizuno, T, Chou, M-Y, and Inouye, M (1984). A unique mechanism regulating gene expression: Translational inhibition by a complementary RNA transcript (micRNA). Proc. Natl Acad. Sci. 81, 1966-1970.
3. Nordström, K, and Wagner, EG (1994). Kinetic aspects of control of plasmid replication by antisense RNA, Trends Biochem. Sci. 19, 294-300.
4. Gottesman, S (2004). The small RNA regulators of Escherichia coli: Roles and mechanisms. Ann. Rev. Microbiol. 58, 303-328.
5. Lee, RC, Feinbaum, RL and Ambros, V (1993). The C. elegans heterochronic gene lin-4 encodes small RNAs with antisense comlementarity to lin-14. Cell 75, 843-854.
6. Wightman, B, Ha, I and Ruvkun, G (1993). Posttranscriptional regulation of the heterochronic gene lin-14 by lin-4 mediates temporal pattern formation in C. elegans. Cell 75, 855-862.
7. Reinhart, BJ, Slack, FJ, Basson, M, Pasquinelli, AE, Bettinger, JC, Rougvie, HR, Horvitz, HR and Ruvkun, G (2000). The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 403, 901-906.
8. Lee, RC and Ambros, V (2001). An extensive class of small RNAs in Caenorhabditis elegans. Science 294, 862-864.
9. Lau, NC, Lim, LP, Weinstein, EG, and Bartel, DP (2001). An abundant class of tiny RNAs with probable regulatory roles in Caenorhabditis elegans. Science 294, 858-862.
10. Lagos-Quintana, M, Rauhut, R, Lendeckel, W and Tuschl, T (2001). Identification of novel genes coding for small expressed RNAs. Science 294, 853-858.
11. Matzke M, Primig, M, Trnovsky, J and Matzke, A (1989). Reversible methylation and inactivation of marker genes in sequentially transformed plants. EMBO J. 8, 643-649.
12. Wassenegger, M, Heimes, S, Riedel, L and Sänger H (1994). RNA-directed de novo methylation of genomic sequences in plants. Cell 76, 567-576.
13. Park, Y-D, Moscone, EA, Iglesis, VA, Vaucheret, H , Matzke, AJM and Matzke, MA (1996). Gene silencing mediated by promoter homology occurs at the level of transcription and results in meiotically heritable alterations in methylation and gene activity. Plant J. 9, 183-194.
14. Napoli, C, Lemieux, C. and Jorgensen, R (1990). Introduction of a chimeric chalcone synthase gene into petunia results in reversible co-suppression of homologous genes in trans. Plant Cell 2, 279-289.
15. van der Krol, A, Mur, L. Beld, M, Mol, JNM, and Stuitje, AR (1990). Flavonoid genes in petunia: Addition of a limited number of gene copies may lead to a suppression of gene expression. Plant Cell 2, 291-299.
16. Smith, CJS, Watson, CF, Bird, CR, Ray, J Schuch, W and Grierson, D (1990). Expression of a truncated tomato polygalacturonase gene inhibits expression of the endogenous gene in transgenic plants. Mol. Gen. Genet. 224, 447-481.
17. de Carvalho, F, Gheyson, G, Kushnir, S. van Montagu, M, Inzé, D et al. (1992). Suppression of beta-1,3-glucanase transgene expression in homozygous plants. EMBO J. 11, 2595-2602.318.
18. van Blokland, K, van der Geest, N, Mol, J, and Kooter, J (1994). Transgene-mediated suppression of chalcone synthase expression in Petunia hybrida results from an increase in RNA turnover. Plant J. 6, 861-877.
19. Romano, N and Macino, G (1992). Quelling: Transient inactivation of gene expression in Neurospora crassa by transformation with homologous sequences. Mol. Microbiol. 6, 3343-3353.
20. Lindbo, J, Silva-Rosales, L, Proebsting, W and Dougherty, W (1993). Induction of a highly specific antiviral state in transgenic plants: Implications for regulation of gene expression and virus resistance. Plant Cell 5, 1749-1759.
21. Dougherty, WG and Parks, TD (1995). Transgenes and suppression: telling us something new? Curr. Opin. Cell Biol. 7, 399-405.
22. Fire, A, Xu, S, Montgomery, MK, Kostas, SA, Driver, SE, and Mello, CC (1998). Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391, 806-811.
23. Izant, JG and Weintraub, H (1984). Inhibition of thymidine kinase gene expression by anti-sense RNA: A molecular approach to genetic analysis. Cell 36, 1007-1015.
24. Guo, S and Kemphues, KJ (1995). Par-1, a gene required for establishing polarity in C. elegans embryos, encodes a putative Sex/Thr kinase that is asymmetrically distributed. Cell 81, 611-620.
25. Rocheleau, CE, Downs WD, Lin, R, Wittman, C, Bei, Y, Cha, YH, Ali, M, Priess JR and Mello, CC (1997), Wnt signaling and an APC-relatred gene specify endoderm in early C. elegans embryos. Cell 90, 707-716.
26. Montgomery, MK, Xu S, and Fire A (1998). RNA as target of double-stranded RNA-mediated genetic interference in Caenorhabditis elegans. Proc. Natl Acad. Sci. 95, 15502-15507.
27. Tuschl, T, Zamore, PD, Lehmann, R, Bartel, DP, and Sharp PA (1999). Targeted mRNA degradation by double-stranded RNA in vitro. Genes Devel. 13, 3191-3197.
28. Elbashir, SM, Harborth, J, Lendeckel, W, Yalcin, A. Weber, K, and Tuschl, T (2001). Duplexes of 21-nucleotide RNAs mediate RNA interference in cultured mammalian cells. Nature 411, 494-498.
29. Hamilton, AJ and Baulcombe, DC (1999). A species of small antisense RNA in posttranscriptional gene silencing in plants. Science 286, 950-952.
30. Waterhouse, PM, Graham, MW, and Wang, M-B (1998). Virusa resistance and gene silencing in plats can be induced by simultaneous expression of sense and antisense RNA. Proc. Natl Acad. Sci. 95, 13959-13964.
31. Zamore, PD, Tuschl, T, Sharp, PA, and Bartel, DP (2000). RNAi: Double-stranded RNA directs the ATP-dependent cleavage of mRNA at 21 to 23 nucleotide intervals. Cell 101, 25-33.
32. Parrish, S, Fleenor, J, Xu S, Mello, C and Fire A (2000). Functional anatomy of a dsRNA trigger: Differential requirements for the two trigger strands in RNA interference. Molec. Cell 6, 1077-1087.
33. Hammond, SM, Bernstein, E, Beach, D, and Hannon, GJ (2000). An RNA-directed nuclease mediates post-transcriptional gene silencing in Drosophila cells. Nature 404, 293-296.
34. Bernstein, E, Caudy, AA, Hammond, SM, and Hannon, GJ (2001). Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 409, 363-366.
35. Cogoni C and Macino G (1999). Gene silencing in Neurospora crassa requires a protein homologus to RNA-depednet RNA polymerase. Nature 399, 166-169.
36. Hannon, GJ (2002), RNA interference. Nature 418, 244-251.
37. Mello, CC and Conte Jr, D (2004). Revealing the world of RNA interference. Nature 431, 338-342.
38. Meister, G and Tuschl, T (2004). Mechanisms of gene silencing by double-stranded RNA. Nature 431, 343-349.
39. Hammond, SM (2005). Dicing and slicing. The core machinery of the RNA interference pathway. FEBS Lett. 579, 5822-5829.
40. Covey, S, Al-Kaff, N. Lángara, A and Turner, D (1997). Plants combat infection by gene silencing. Nature 385, 781-782.
41. Ratcliff, F, Harrison, B and Baulcombe, D (1997). A similarity between viral defence and gene silencing in plants. Science 276, 1558-1560.
42. Ketting, RF, Haverkamp, TH, van Luenen HG, and Plasterk, RH (1999). Mut-7 of C. elegans, required for transposon silencing and RNA interference, is a homolog of Werner syndrome helicase and RNase D. Cell 99, 133-141.
43. Tabara, H, Sarkissian M, Kelly WG, Fleenor, J, Grishok, A, Timmons, L, Fire, A and Mello CC (1999). The rde-1 gene, RNA interference, and transposon silencing in C. elegans. Cell 99, 123-132.
44. Plasterk, RHA (2002). RNA Silencing: The Genome’s Immune System. Science 296, 1263-1265.
45. Reinhart, BJ, Weinstein, EG, Rhodes, MW, Bartel, B and Bartel DP (2002). MicroRNAs in plants. Genes Dev. 16, 1616-1626.
46. Sharp, PA (2006). The Biology of Short RNAs. In “RNA World”, Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY.
47. Murchison, EP and Hannon, GJ (2004). miRNAs on the move: miRNA biogenesis and the RNAi machinery. Curr. Opin. Cell Biol. 16, 223-229.
48. Zamore, PD and Haley, B (2005). Ribo-gnome: The Big World of Small RNAs. Science 309, 1519-1524.
49. Mette, MF, Aufsatz, W, van der Winden, J, Matzke, M and Matzke A (2000). Transcriptional gene silencing and promoter methylation triggered by double stranded RNA. EMBO J. 19, 5194-5201.
50. Sijen, T, Vijn, I, Rebocho, A, van Blokland R, Roelofs, D et al. (2001). Transcriptional and posttranscriptional gene silencing are mechanistically related. Curr. Biol 11, 436-440.
51. Hall, IM, Shankaranarayana, GD, Noma K, Ayoub, N, Cohen, A and Grewal SI (2002). Establishment and maintenance of a heterochromatic domain. Science 297, 2232-2237.
52. Volpe, TA, Kidner, C, Hall, IM, Teng, G, Grewal, SI and Martienssen, RA (2002). Regulation of heterochromatic silencing and histone H3 lysine-9 methylation by RNAi. Science 297, 1833-1837.
53. Dorsett, Y and Tuschl, T (2004). siRNAs: Applications in functional genomics and potential as therapeutics. Nature Reviews 3, 318-329.
54. Hannon, GJ and Rose, JJ (2004). Unlocking the potential of the human genome with RNA interference. Nature 431, 371-378.
55. Soutschek, J et al. (2004). Therapeutic silencing of an endogenous gene by systematic administration of modified siRNAs. Nature 432, 173-178.
56. Morrisey, DV et al. (2005). Potent and persistent in vivo anti-HBV activity of chemically modified siRNAs. Nat. Biotechnol. 23, 1002-1007.
57. Palliser, D (2006). An siRNA-based microbicide protects mice from lethal herpes simplex virus infection. Nature 439, 89-94.
58. Zimmermann TS et al. (2006). RNAi-mediated gene silencing in non-human primates. Nature 441, 111-114.
Acknowledgements.
I am grateful to Annika Röhl for designing the figures and to Adam Smith for helpful comments on the text.
Bertil Daneholt
Professor of Molecular Genetics, Karolinska Institutet, Stockholm
Chairman of the Nobel Assembly
Nobel Prizes and laureates
Six prizes were awarded for achievements that have conferred the greatest benefit to humankind. The 14 laureates' work and discoveries range from quantum tunnelling to promoting democratic rights.
See them all presented here.
