Popular information
Popular science background:
A (r)evolution in chemistry (pdf)
Populärvetenskaplig information:
En (r)evolution inom kemin (pdf)

The Nobel Prize in Chemistry 2018
The power of evolution is revealed through the diversity of life. The Nobel Prize in Chemistry 2018 is awarded to Frances H. Arnold, George P. Smith and Sir Gregory P. Winter for the way they have taken control of evolution and used it for the greatest benefit to humankind. Enzymes developed through directed evolution are now used to produce biofuels and pharmaceuticals, among other things. Antibodies evolved using a method called phage display can combat autoimmune diseases and, in some cases, cure metastatic cancer.
A (r)evolution in chemistry
We live on a planet where a powerful force has become established: evolution. Since the first seeds of life appeared around 3.7 billion years ago, almost every crevice on Earth has been filled by organisms adapted to their environment: lichens that can live on bare mountainsides, archaea that thrive in hot springs, scaly reptiles equipped for dry deserts and jellyfish that glow in the dark of the deep oceans.
In school, we learn about these organisms in biology, but let’s change perspective and put on a chemist’s glasses. Life on Earth exists because evolution has solved numerous complex chemical problems. All organisms are able to extract materials and energy from their own environmental niche and use them to build the unique chemical creation that they comprise. Fish can swim in the polar oceans thanks to antifreeze proteins in their blood and mussels can stick to rocks because they have developed an underwater molecular glue, to give just a few of the innumerable examples.
The brilliance of life’s chemistry is that it is programmed into our genes, allowing it to be inherited and developed. Small random changes in genes change this chemistry. Sometimes this leads to aweaker organism, sometimes a more robust one. New chemistry has gradually developed and life on Earth has become increasingly complex.

Figure 1. The Nobel Laureates in Chemistry 2018 have harnessed evolution and further advanced it in their laboratories.
Figure 1. The Nobel Laureates in Chemistr y 2018 have harnessed
evolution and further advanced it in their laboratories.
This process has now come so far that it has given rise to three individuals so complex they have managed to master evolution themselves. The Nobel Prize in Chemistry 2018 is awarded to Frances H. Arnold, George P. Smith and Sir Gregory P. Winter, because they have revolutionised both chemistry and the development of new pharmaceuticals through directed evolution. Let’s begin with the star of enzyme engineering: Frances Arnold.
Enzymes – the sharpest chemical tools of life
Even in 1979, as a newly graduated mechanical and aerospace engineer, Frances Arnold had a clear vision: to benefit humanity through the development of new technology. The US had decided that 20 per cent of its power would come from renewable sources by 2000, and she worked with solar power. However, prospects for the future of this industry changed radically after the 1981 presidential election, so instead she turned her gaze to the new DNA technology. As she expressed it herself: “It was clear that a whole new way of making materials and chemicals that we needed in our daily lives, would be enabled by the ability to rewrite the code of life.”
Instead of producing pharmaceuticals, plastics and other chemicals using traditional chemistry, which often requires strong solvents, heavy metals and corrosive acids, her idea was to use the chemical tools of life: enzymes. They catalyse the chemical reactions that occur in the Earth’s organisms and, if she learned to design new enzymes, she could fundamentally change chemistry.
Human thought has limitations
Initially, like many other researchers at the end of the 1980s, Frances Arnold attempted to use a rational approach to rebuilding enzymes to give them new properties, but enzymes are extremely complex molecules. They are built from 20 different kinds of building blocks – amino acids – that can be infinitely combined. A single enzyme can consist of several thousand amino acids. These are linked together in long chains, which fold up into special three-dimensional structures. The environment necessary to catalyse a particular chemical reaction is created inside these structures.
Using logic to try to work out how this elaborate architecture should be remodelled to give an enzyme new properties is difficult, even with contemporary knowledge and computer power. In the early 1990s, humble in the face of nature’s superiority, Frances Arnold decided to abandon this, in her words, “somewhat arrogant approach” and instead found inspiration in nature’s own method for optimising chemistry: evolution.
Arnold starts to play with evolution
For several years, she had tried to change an enzyme called subtilisin so that rather than catalysing chemical reactions in a water-based solution, it would work in an organic solvent, dimethylformamide (DMF). Now she created random changes – mutations – in the enzyme’s genetic code and then introduced these mutated genes into bacteria that produced thousands of different variants of subtilisin.
After this, the challenge was to find out which of all these variants worked best in the organic solvent. In evolution, we talk about survival of the fittest; in directed evolution this stage is called selection.
Frances Arnold utilised the fact that subtilisin breaks down milk protein, casein. She then selected the variant of subtilisin that was most effective in breaking down casein in a solution with 35 per cent DMF. She subsequently introduced a new round of random mutations in this subtilisin, which yielded a variant that worked even better in DMF.
In the third generation of subtilisin she found a variant that worked 256 times better in DMF than the original enzyme. This variant of the enzyme had a combination of ten different mutations, the benefits of which no one could have worked out in advance.
With this, Frances Arnold demonstrated the power of allowing chance and directed selection, instead of solely human rationality, to govern the development of new enzymes. This was the first and most decisive step towards the revolution we are now witnessing.
The next important step was taken by Willem P. C. Stemmer, a Dutch researcher and entrepreneur who died in 2013. He introduced yet another dimension to the directed evolution of enzymes: mating in a test tube.
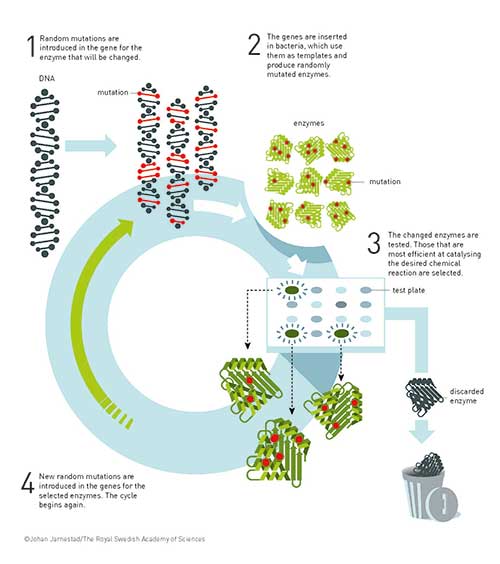
Figure 2. The underlying principle for the directed evolution of enzymes. After a few cycles of directed evolution, an enzyme may be several thousand times more effective.
Figure 2. The underlying principle for the directed evolution of enzymes. After a few cycles of directed evolution, an enzyme may be several thousand times more effective.
Mating – for more stable evolution
One prerequisite for natural evolution is that genes from different individuals are mixed through mating or pollination, for example. Beneficial properties can then be combined and give rise to a more robust organism. At the same time, less functional gene mutations can disappear from one generation to another.
Willem Stemmer used the test tube equivalent to mating: DNA shuffling. In 1994 he demonstrated that it was possible to cut different versions of a gene into small pieces and then, helped by the tools of DNA technology, puzzle together the pieces into a complete gene, one that is a mosaic of the original versions.
Using several cycles of DNA shuffling, Willem Stemmer changed an enzyme so that it became much more effective than the original enzyme. This showed that mating genes together – researchers call this recombination – can result in the even more efficient evolution of enzymes.
New enzymes produce sustainable biofuel
The tools of DNA technology have been refined since the early 1990s, and the methods used in directed evolution have multiplied. Frances Arnold has been at the leading edge of these developments; the enzymes now produced in her laboratory can catalyse chemistry that does not even exist in nature, producing entirely new materials. Her tailored enzymes have also become important tools in the manufacture of various substances, such as pharmaceuticals. Chemical reactions are sped up, produce fewer by-products and, in some cases, it has been possible to exclude the heavy metals required by traditional chemistry, thus considerably reducing environmental impact.
Things have also come full circle: Frances Arnold is again working with the production of renewable energy. Her research group has developed enzymes that transform simple sugars to isobutanol, an energy-rich substance that can be used for the production of both biofuels and greener plastics. One long-term aim is to produce fuels for a more environmentally friendly transport sector. Alternative fuels – produced by Arnold’s proteins – can be used in cars and aeroplanes. In this way, her enzymes are contributing to a greener world.
And now to the second half of 2018’s Nobel Prize in Chemistry, where directed evolution has instead led to pharmaceuticals that can neutralise toxins, combat the progression of autoimmune diseases and, in some cases, even cure metastatic cancer. This is where a vital role is played by a tiny virus that infects bacteria and the method known as phage display.
Smith uses bacteriophages
As is so often the case, science took an unpredictable path. In the first half of the 1980s, when George Smith started to use bacteriophages – viruses that infect bacteria – it was mainly in the hope that they could be used to clone genes. DNA technology was still young and the human genome was like an undiscovered continent. Researchers knew that it contained all the genes required to produce the body’s proteins, but identifying the specific gene for a particular protein was more difficult than looking for a needle in a haystack.
However, there were huge benefits for the scientist who did find it. Using that time’s new genetic tools, the gene could be inserted into bacteria which – with a bit of luck – could mass produce the protein to be studied. The whole process was called gene cloning and George Smith’s idea was that researchers looking for genes could use bacteriophages in an ingenious way.
Bacteriophages – a link between a protein and its unknown gene
Bacteriophages are simple by nature. They consist of a small piece of genetic material that is encapsulated in protective proteins. When they reproduce, they inject their genetic material into bacteria and hijack their metabolism. The bacteria then produce new copies of the phage’s genetic material and the proteins that form the capsule, which form new phages.
George Smith’s idea was that researchers should be able to use the phages’ simple construction to find an unknown gene for a known protein. At this time, large molecular libraries were available, which contained masses of fragments of various unknown genes. His idea was that these unknown gene fragments could be put together with the gene for one of the proteins in the phage capsule.
When new phages were produced, the proteins from the unknown gene would end up on the surface of the phage as part of the capsule protein (figure 3).
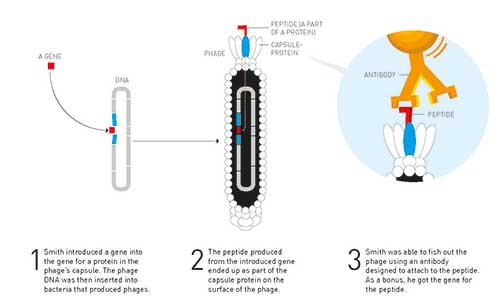
Figure 3. Phage display – George Smith developed this method for finding unknowns genes for a known protein
Figure 3. Phage display – George Smith developed this method for finding unknowns genes for a known protein
Antibodies can fish out the right protein
This would result in a mixture of phages that carried multitudes of different proteins on their surface. In the next stage – George Smith postulated – researchers would be able to use antibodies to fish phages carrying various known proteins out of this soup. Antibodies are proteins that function like targeted missiles; they can identify and bind to a single protein among tens of thousands of others with extreme precision. If researchers caught something in the phage soup using an antibody that they knew attached to a known protein, as a bycatch they would get the thus-far unknown gene for the protein.
It was an elegant idea and, in 1985, George Smith demonstrated that it could work. He produced a phage that carried part of a protein, a peptide, on its surface. Using an antibody, he was then able to fish the phage he had constructed out of a soup of many phages.
Through this experiment, George Smith laid the foundation of what has come to be known as phage display. The method is brilliant in its simplicity. Its strength is that the phage functions as a link between a protein and its gene. However, it was not within gene cloning that the method had its major breakthrough; instead, in around 1990, several research groups started to use phage display to develop new biomolecules. One of the people who adopted the technique was Gregory (Greg) Winter and it is thanks to his research that phage display is now bringing great benefit to mankind. To understand why, we need to take a closer look at antibodies.
Antibodies can block disease processes
The human lymphatic system has cells that can produce hundreds of thousands of different kinds of antibodies. Using a well-developed system, all these cells have been tested so that no antibody attaches to any of the body’s various types of molecules. However, this enormous variation ensures that there is always an antibody that attaches to the viruses or bacteria that infect us. When an antibody attaches to them it sends a signal to aggressive immune cells to destroy the invaders.
Because antibodies are highly selective and can attach themselves to a single molecule among tens of thousands of others, researchers had long hoped it would be possible to design antibodies that block various disease processes in the body and function like pharmaceuticals. Initially, to obtain these therapeutic antibodies, mice were injected with different targets for pharmaceuticals, such as proteins from cancer cells. However, in the 1980s it became increasingly clear that this method had limitations; some substances were toxic for the mice and others did not result in any antibody production. Additionally, it was discovered that the obtained antibodies were recognised as foreign by the patients’ immune system, which attacked them. This led to the mice antibodies being destroyed and there was a risk of side effects for the patients.
It was this obstacle that caused Greg Winter to start investigating the potential of George Smith’s phage display. He wanted to avoid using mice and to be able to base pharmaceuticals on human antibodies because they are tolerated by our immune system.
Winter puts antibodies on the surface of phages
Antibodies are Y-shaped molecules; it is the far end of each arm that attaches to foreign substances. Greg Winter joined the genetic information for this part of the antibody to the gene for one of the phage’s capsule proteins and, in 1990, he demonstrated that this led to the antibody’s binding site ending up on the surface of the phage. The antibody he used was designed to attach to a small molecule known as phOx. When Greg Winter used phOx as a kind of molecular fishing hook, he succeeded in pulling the phage with the antibody on its surface out of a soup of four million other phages.
After this, Greg Winter showed that he could use phage display in the directed evolution of antibodies. He built up a library of phages with billions of varieties of antibodies on their surfaces. From this library, he fished out antibodies that attached to different target proteins. He then randomly changed this first generation of antibodies and created a new library, in which he found antibodies with even stronger attachments to the target. For instance, in 1994 he used this method to develop antibodies that attached to cancer cells with a high level of specificity.
The world’s first pharmaceutical based on a human antibody
Greg Winter and his colleagues founded a company based on the phage display of antibodies. In the 1990s, it developed a pharmaceutical entirely based on a human antibody: adalimumab. The antibody neutralises a protein, TNF-alpha, that drives inflammation in many autoimmune diseases. In 2002, the pharmaceutical was approved for the treatment of rheumatoid arthritis and is now also used for treating different types of psoriasis and inflammatory bowel diseases.
The success of adalimumab has spurred significant development in the pharmaceutical industry and phage display has been used to produce cancer antibodies, amongst others. One of these releases the body’s killer cells so they can attack tumour cells. Tumour growth slows down and, in some cases, even patients with metastatic cancer are cured, which is a historic breakthrough in cancer care.
Another antibody pharmaceutical that has been approved neutralises the bacterial toxin that causes anthrax, while another slows the autoimmune disease known as lupus; many more antibodies are currently undergoing clinical trials, for example to combat Alzheimer’s disease.
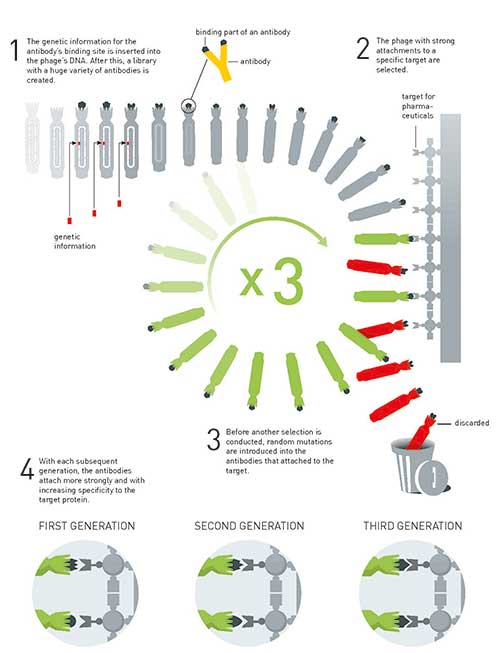
Figure 4. The principle for the directed evolution of antibodies using phage display. This method is used to produce new pharmaceuticals.
Figure 4. The principle for the directed evolution of antibodies using phage display. This method is used to produce new pharmaceuticals.
The start of a new era in chemistry
The methods that the 2018 Nobel Laureates in Chemistry have developed are now being internationally developed to promote a greener chemicals industry, produce new materials, manufacture sustainable biofuels, mitigate disease and save lives. The directed evolution of enzymes and the phage display of antibodies have allowed Frances Arnold, George Smith and Greg Winter to bring the greatest benefit to humankind and to lay the foundation for a revolution in chemistry.
Links and further reading
Additional information on this year’s prizes, including a scientific background in English, is available on the website of the Royal Swedish Academy of Sciences, www.kva.se, and at http://nobelprize.org. There you can watch video footage of the press conferences, the Nobel Lectures and more. Information on exhibitions and activities related to the Nobel Prizes and the Prize in Economic Sciences is available at www.nobelcenter.se.
Articles
Arnold, F. & Macuare, K. A. (2016) The NAI Fellow Profile: An Interview with Dr. Frances Arnold. Technology and Innovation, 18, 79-82
Harding, A. (2006) Profile Sir Greg Winter—humaniser of antibodies. Lancet, 368, S50
Nightingale, K. (2013) Greg Winter: Pioneering antibody drugs. Insight, www.insight.mrc.ac.uk/2013/02/27/greg-winter-pioneering-antibody-drugs/
Trager, R. (2018) In situ with Frances Arnold. Chemistry World, www.chemistryworld.com/culture/in-situ-with-frances-arnold/3008732.article
Videos
MoleCluesTV (4 June 2017) Frances Arnold: New enzymes by evolution
www.youtube.com/watch?v=05r-FLGtsEQ
SlidesLive (23 Nov 2015) Therapeutics Antibodies: A Revolution in Pharmaceuticals
The Royal Swedish Academy of Sciences has decided to award the Nobel Prize in Chemistry 2018
with one half to
FRANCES H. ARNOLD
Born 1956 in Pittsburgh, USA. Ph.D. 1985, University of California, Berkeley, USA. Linus Pauling Professor of Chemical Engineering, Bioengineering and Biochemistry, California Institute of Technology, Pasadena, USA.
http://fhalab.caltech.edu
“for the directed evolution of enzymes”
and the other half jointly to
GEORGE P. SMITH
Born 1941 in Norwalk, USA. Ph.D. 1970, Harvard University, Cambridge, USA. Curators’ Distinguished Professor Emeritus of Biological Sciences, University of Missouri, Columbia, USA.
http://biology.missouri.edu/people/?person=94
and
SIR GREGORY P. WINTER
Born 1951 in Leicester, UK. Ph.D. 1976. University of Cambridge, UK. Research Leader Emeritus, MRC Laboratory of Molecular Biology, Cambridge, UK.
www2.mrc-lmb.cam.ac.uk/group-leaders/emeritus/greg-winter/
“for the phage display of peptides and antibodies”
Science Editors: Claes Gustafsson, Gunnar von Heijne and Sara Snogerup Linse, the Nobel Committee for Chemistry
Text: Ann Fernholm
Translation: Clare Barnes
Illustrations: © Johan Jarnestad/The Royal Swedish Academy of Sciences
Editor: Ann Fernholm
© The Royal Swedish Academy of Sciences
Nobel Prizes and laureates
Six prizes were awarded for achievements that have conferred the greatest benefit to humankind. The 14 laureates' work and discoveries range from quantum tunnelling to promoting democratic rights.
See them all presented here.
