Popular information
Information for the public:
Central to life (pdf)
Populärvetenskaplig information:
Nödvändigt för allt liv (pdf)

The Nobel Prize in Chemistry 2006
This year’s Nobel Prize in Chemistry is awarded to Roger D. Kornberg for his fundamental studies concerning how the information stored in the genes is copied, and then transferred to those parts of the cells that produce proteins. Kornberg was the first to create an actual picture of this process at the molecular level, in the important group of organisms called eukaryotes (which, as opposed to bacteria, have well-defined cell nuclei). Mammals like ourselves, as well as ordinary yeast, belong to this group of organisms.
Central to life
Constant transcription of the genetic information in the DNA is a central process for all living beings. The DNA-molecule itself lies protected within the cell nuclei. The genetic information therefore needs to be copied and transformed into what is called messenger-RNA, which carries the information out into the protein-producing parts of the cells. It is the proteins that in their turn actually construct the organism and its function.
If the transcription process is interrupted the organism will soon die, since all proteinproduction in the cells ceases. This is what happens in cases of poisoning by certain types of toadstools, like the death cap. The toxin in the death cap blocks the function of a specific enzyme, RNA-polymerase, which plays a central role in the transcription process. Within a few days the toxin spreads from the bowels to the liver and kidneys, hence slowly destroying all these organs.
Many illnesses – like cancer, heart disease, and different kinds of inflammation – are also linked to disturbances in the transcription process.
Explains variety
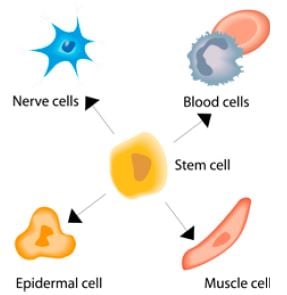
It is thanks to the flexible transcription system in eukaryotes that such complexity is possible within this group of organisms. All the cells of the body contain the same genetic information, but there is great variation between organs in the information that is actually transcribed, and thereby translated into protein production. Knowledge about the transcription process is also fundamental for understanding how stem cells develop into different kinds of specific cells, with well-defined functions in different organs. The interest in stem cells for therapeutic applications is very largely based on their capacity to develop into any kind of functional cell in a living organism. Understanding more about how transcription is regulated is therefore one of the necessary steps, if we want to realise the full potential of stem cells in medicine.
© The Royal Swedish Academy of Sciences
The DNA-molecule is constructed using four different types of building blocks: G, C, A and T. The RNA is composed of four corresponding types of building blocks. The genetic information written into these molecules is determined by the sequence in which the building blocks are put together. The genetic code therefore consists of an alphabet with only four letters.
The DNA-molecule is a double helix, with a G on one of the strands always opposing a C on the other strand, and A in a similar manner opposing T (Watson, Crick and Wilkins were awarded their famous Nobel Prize in Physiology or Medicine in 1962 for this discovery). The transcription process starts with the DNA-double helix opening up so that a strand of “naked” building blocks can be used as a template in creating an RNA-strand. The cell contains the RNA building blocks in solution. If the transcription process functions, the RNA building block corresponding to a G inserts itself opposite a C on the DNA-strand, the RNA-version of a T inserts itself opposite an A on the DNA-strand, and vice versa. In this manner, the RNA-strand is constructed in a step-wise process to form a “negative” of the DNA-strand. One fundamental question is how this process works in more detail. The wellbeing of the organism depends on translation of the correct information into proteins. Not more than one fault in ten thousand can be tolerated before the organism is impaired. For this reason the mechanism ensuring that the building blocks of RNA are put together in the right order must be very specific. The key lies in a special enzyme that carefully governs this process: RNA-polymerase.
A snapshot of life
The picture below showing RNA-polymerase in full action was created by Kornberg as recently as 2001. The big white molecule (which looks like a bundle of wires) is the RNA-polymerase, serving as a support for the DNA-strand (in blue). The polymerase molecule keeps the DNA-strand in the right position during transcription and creates a minuscule “cavity”, so small that it will only accept the RNA building block that fits with the opposing DNA building block exactly. If the wrong RNA building block tries to insert itself into the cavity it simply does not fit, just as the wrong piece cannot be inserted in a half completed jigsaw puzzle. This leads to the step-wise construction of the RNA-strand (in red). Once a new building block has been inserted in the right place, the DNA-strand is prompted forward by a small helical structure (in green) in the polymerase. This spring-like structure flips back and forth thanks to constant spontaneous changes in shape of the polymerase (this is precisely the mechanism which is destroyed by the toxin of the death cap, mentioned above). In this manner, the DNA-strand is again and again moved into the right position for a new building block to be added to the growing RNA-strand.
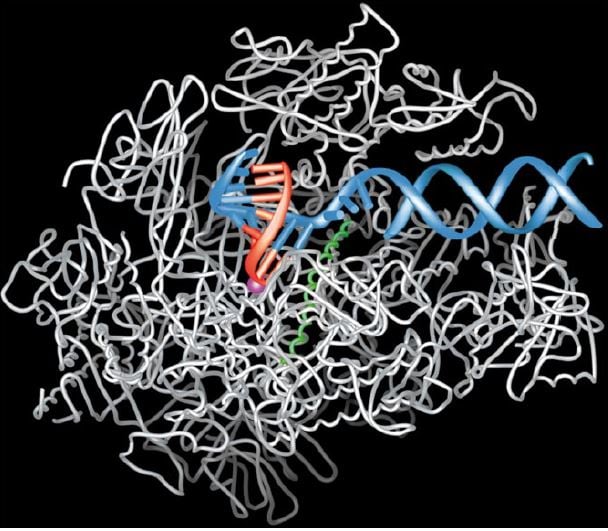
Figure 1. The transcription process as depicted by Roger Kornberg in 2001. RNA-polymerase in white, DNA-helix in blue and the growing RNA-strand in red.
© The Royal Swedish Academy of Sciences
The truly revolutionary aspect of the picture Kornberg has created is that it captures the process of transcription in full flow. What we see is an RNA-strand being constructed, and hence the exact positions of the DNA, polymerase and RNA during this process. In an ingenious manner Kornberg has managed to freeze the construction process of RNA half-way through, simply by leaving out one of the necessary building blocks from the solution he uses. When the construction reaches the point where the missing building block should be inserted the whole process stops. Kornberg has then created crystals of the molecules involved, and “taken their picture” using X-rays. This picture is of course not an ordinary photo but an image of the molecules in their crystalline form. Starting with such a picture, a computer can calculate the real position of atoms in the molecules. The image shown here, has thus been created by a computer.
This method of making crystals of biological molecules in order to depict them is a very common tool today. Normally, however, we only see pictures of finished complexes and individual molecules. To capture a chemical reaction as it takes place is very difficult, and it is not enough to be a good crystallographer in order to succeed. What is particular about the way Kornberg works is that he combines crystallography with very deep biochemical knowledge which makes it possible for him to assume full control of the process he wants to depict. Besides this detailed picture of the role of RNA-polymerase, he has also contributed several other important pieces of information concerning the transcription process by studying other crystals of RNA-polymerase, DNA, RNA and important complexes called general transcription factors. These images make it possible to understand the molecular mechanisms governing the transcription process.
Bacteria, mammals and yeast
For a long time it was believed that the transcription process in eukaryotes was very similar to that in bacteria (which unlike eukaryotic cells lack a well-defined nucleus). It has however become clear that the process in eukaryotes is much more complex. RNA-polymerase was actually first found in mammalian liver cells, but it proved very difficult to work with these cells. Instead, bacteria became the first organisms to be studied in detail, since they were much easier to use.
In 1965 the Nobel Prize in Physiology or Medicine was awarded to Jacques Monod, André Lwoff and François Jacob, among other things for their description of how transcription in bacteria works. Besides RNA-polymerase another molecule called sigma factor is necessary in order for transcription in bacteria to start. This molecules binds to RNA-polymerase and, through its capacity to recognize special codes in the DNA, it tells where the genetic information starts and where it ends. Without the sigma factor, transcription would never start since the polymerase on its own does not “know” where on the DNA-strand to start reading.
However, when researchers returned to eukaryotic cells after this discovery no one managed to find the sigma factor. It gradually transpired that in eukaryotes there are five different molecular complexes which correspond to the single sigma factor in bacteria. All of them are necessary for transcription to start in our cells. These complexes are called general transcription factors and play an important role in the fine-tuning of transcription in eukaryotes. Finding all these five transcription factors required a great deal of painstaking work, purifying cell extracts in several steps. By removing one substance at a time from these extracts until the transcription stopped working, researchers isolated the substances that were necessary for transcription to take place. When this was done, many were convinced that every part of the transcription system in eukaryotes had been discovered, just as with bacteria. Even so, the great variety in shape and function of eukaryotic cells had not been explained. There were as yet no good answers to questions concerning how certain genes are expressed in blood cells, others in the cells of the liver, and so on.
One of Kornberg’s great contributions at this stage was his development of a new system of using yeast cells in laboratory work. Ordinary yeast is a eukaryote, just as mammals are. This makes it possible to use yeast as a model organism instead of cells from mammal organs. Yeast cells are much easier to manipulate and it is easier to create a homogenous material to work with using these cells. In spite of this, Kornberg’s research group spent ten years fine-tuning the system, before it could be used to investigate the transcription process. Many research groups would have given up much earlier, since several years went by without any actual results that could be published. But it is this very system, using yeast cells, that has made it possible for Kornberg to produce RNA-polymerase and general transcription factors in the adequate form and quantity to create crystals to investigate.
The necessary relay
Thanks to this system of yeast cells, Kornberg found yet another molecular complex which turned out to play an important role as a kind of on-off switch for the transcription process in eukaryotes. The DNA-helix includes parts called enhancers which bind to specific substances that are found in different tissues. In this manner transcription of certain genes is stimulated in certain tissues. In the liver – for instance – there is a specific signalling substance, which binds to a liver-enhancer in the DNA hence turning on the transcription of a gene close to the enhancer. In other parts of the body, this liver-specific gene will however never be switched on because the necessary signalling substance is lacking. Kornberg discovered that this regulation requires the presence of yet another molecular complex, transmitting the signals and thereby switching the transcription on or off. This “relay”-complex is called Mediator.
The great complexity of eukaryotic organisms is actually enabled by the fine interplay between tissue-specific substances, enhancers in the DNA and Mediator. The discovery of Mediator is therefore a true milestone in the understanding of the transcription process.
To be continued
Kornberg continues his crystallographic and functional work today, by trying to include ever larger parts of the transcription apparatus in his descriptions. Hitherto he has shown the interplay between RNA-polymerase, DNA and the growing RNA-strand and its building blocks. The next step is to include all the general transcription factors and especially Mediator in full action. Kornberg has already taken the first steps in this direction.
Crystallography is a particularly important tool in this context because the actual spatial configuration of the different components in the transcription system plays such a decisive role. It is very difficult to create an understanding of the process through traditional chemical methods, which do not tell us anything about the spatial positions of the molecules and atoms. Many of the components in the process undergo no important chemical changes, we need to quite physically “see” the molecules and their positions at different stages in the process to understand how the transcription works.
The gradual construction of a functional image of how transcription works ultimately involves attaining understanding of how the genetic information generates the great variety of living organisms around us. Knowledge about how this happens in the human body is also of fundamental medical importance.
Links and further reading
The Academy’s website, www.kva.se, and http://nobelprize.org have more information on this year’s Prizes, including a web-TV broadcast of the press conference and advanced information mainly intended for the research community.
Original scientific articles
Cramer, P., Bushnell, D.A. and Kornberg, R.D. (2001) Structural basis of transcription: RNA polymerase II at 2.8 ångstrom resolution. Science 292, 1863-1876.
Gnatt, A.L., Cramer, P., Fu, J., Bushnell, D.A. and Kornberg, R.D. (2001) Structural basis of transcription: An RNA polymerase II elongation complex at 3.3 Å resolution. Science 292, 1876-1882.
Bushnell, D.A., Westover, K.D., Davis, R.E. and Kornberg, R.D. (2004) Structural basis of transcription: An RNA polymerase II – TFIIB cocrystal at 4.5 angstroms. Science 303, 983-988.
Review article
Boeger, H., Bushnell, D.A., Davis, R., Griesenbeck, J., Lorch, Y., Strattan, J.S., Westover, K.D. and Kornberg, R.D. (2005). Structural basis of eukaryotic gene transcription. FEBS Lett. 579, 899-903.
Link
Film of transcription: The Dolan DNA learning center – genes in education. http://www.dnalc.org/home.html. Media showcase; Transcription: DNA codes for mRNA, 3D animation.
THE LAUREATE
ROGER D. KORNBERG
Stanford University School of Medicine, Fairchild building, 1st floor
299 Campus Dr. Stanford, California 94 305-5126 USA
http://kornberg.stanford.edu/
US citizen. Born 1947 (59) in St Louis, MO, USA. PhD from Stanford University, CA, USA. Mrs. George A. Winzer Professor in Medicine at Stanford University School of Medicine, CA, USA.
© The Royal Swedish Academy of Sciences
Nobel Prizes and laureates
Six prizes were awarded for achievements that have conferred the greatest benefit to humankind. The 14 laureates' work and discoveries range from quantum tunnelling to promoting democratic rights.
See them all presented here.
