Bruce A. Beutler
Biographical

One of the happiest days of my life
It was 2:30 a.m. on October 3, 2011, and I was at home, in my small condominium in San Diego, CA. I was sleepless, having recently returned from a trip to Hong Kong. There I had shared the Shaw Prize in Life Sciences: one of a series of prizes in recent years. The previous afternoon, as we visited our mother in La Jolla, my brother Earl had remarked that the Nobel Prize in Physiology or Medicine would be announced the next day. He also had asked if I was going to win it this year. This was on my mind as I lay awake.
Bleary-eyed, I looked at my cell phone to see if there was any email. There was a message: just one. The title line seemed to be “Nobel Prize.” I reached for my glasses, looked again, and found I was correct. Nobel Prize. Perhaps this year the Committee had made a mass e-mailing of the announcement to members of various National Academies? I opened the message and read the first lines of a letter from Göran Hansson …
“Dear Dr Beutler,
I have good news for you. The Nobel Assembly has today decided to award you the Nobel Prize in Physiology or Medicine for 2011. You will share the Prize with Drs Jules Hoffmann and Ralph Steinman. Congratulations!”
I was too excited to read more at that moment, and hurried downstairs. I called Betsy Layton, my longtime colleague and Administrative Manager. She was in Dallas, preparing the way for our relocation from the Scripps Research Institute to UT Southwestern, where we had earlier done some of our most important work. She had worked with me over the past 25 years, and had played an important part in finding the Lps mutation for which the Prize was awarded. I woke her from a sound sleep. “Bets,” I said tentatively, “I think I won the Nobel Prize!” She was ecstatic, but I cautioned her that I had to confirm it, and kept her on the line.
I tried to access the website Nobelprize.org, but found I couldn’t; there was too much traffic. Then I went to news.google.com. I searched for my name, and within a minute or two, began to see it in reports emanating from all over the world. It was true. I told Betsy so, and she was so happy she began to cry! Yet still I was somehow disbelieving and only over the days and weeks that followed did the new reality settle in my mind.
Nadia Krochin, my dear friend and companion of many years, called me from the east coast before I could call her, and left a message on my voice mail that was almost incoherent with joy! I called my brother Earl; then my brother Steve; then my sons; then my friend Ari Theofilopoulos. I waited until about 4 a.m. to call my mother. Each person, though awakened abruptly, was nearly as happy as I had been.
By this time, my cell phone was ringing almost constantly, and it didn’t stop ringing until late afternoon. Mostly reporters were calling, and I attempted to respond immediately to a few of them, hearing a constant beeping of incoming calls in the background as I did. Among those to whom I spoke during those first hours was a National Public Radio anchor, and it was on NPR, driving to work in various parts of the country, that many of my friends soon heard the news, along with my initial reaction. Emails were arriving every few seconds, and by the end of the day, I had more than a thousand of them. I realized it was hopeless to reply to them, and instead, via Betsy, I advanced my pre-existing plans to fly to Dallas. While waiting for my 10 a.m. flight, I went for a haircut earlier scheduled for 8 a.m. Hannah Andrusky, a charming lady who had cut my hair each month for several years, agreed to meet me an hour early that morning. I was her first client of the day, and she, too, knew of the Prize. She was as delighted and proud of me as everyone else had been. The haircut was free.
In Dallas I was greeted as a hero. A press conference was arranged for the next day, and after introductory speeches by the Dean, President, and Past President of UT Southwestern Medical Center, I stepped to the podium, met by a warm standing ovation. I spoke for several minutes from the heart, without slides, notes, or preparation of any kind. I felt a sense of self-confidence I hadn’t known before. After all, I was a Nobel Laureate. Everyone present was elated and wished me well. I talked a bit about my life, about the steps that had brought me to the Nobel Prize, and also about some of the struggles and strains this entailed.
My background
All of my grandparents immigrated to the United States to escape persecution as Jews, and I was reminded of this often from an early age. World War II had ended only 12 years before I was born, and recollections were fresh. My father was born in Germany, my mother in America to immigrant parents. My siblings, my cousins and I were always strongly conscious of our European origins, and of the extreme anti-Semitism that had recently prevailed in Europe. Probably we all felt a need to excel partly because of these facts; to show that we were as good as the other children in our schools.
My mother’s parents, Aral (“Harry”) Fleisher (1895–1953), and Miriam (“Mary”) Fleisher, née Krasne (1893–1966), were both from Kiev, and moved separately to the USA near the turn of the century, settling in Chicago where they met and married in 1922. Harry initially worked in the insurance business. However, he was persuaded to join the company of Mary’s father, Louis Krasne (L. Krasne and Sons). Mary and her brother Isador traveled to the west, ultimately settling in Tulsa, Oklahoma where they established a branch of L. Krasne and Sons that traded in “Leather and Findings.” My mother, Brondelle May Fleisher, was born in Chicago but raised in Tulsa from the age of 11. She was the second of three children (Beverly was older and Lois younger), and she survives both her sisters today.
My mother’s father died before I was born, but I knew my maternal grandmother well. Afflicted with a brittle form of diabetes (oddly at a mature age), she lived with us, was known as “Bubbie,” and took care of me during my first few years of life. I remember her as a warm and kindly old woman with a strong Yiddish accent and extremely long hair, nearly touching the ground. Almost blind, she would spend much of her time in the garden, pulling weeds from the lawn, or otherwise trying to be useful. She was happy to talk to me, or walk with me or explore the yard. One of our routines was to “search for the Fountain of Youth” (which always turned out to be a drinking fountain in the back yard). She died within days of having a myocardial infarction at our California home in 1966.
My father’s parents, Alfred Beutler (1891–1962) and Kaethe Beutler née Italiener (1896–1999) were both physicians: he an internist with a particular interest in cardiology, and she a pediatrician. They emigrated from Berlin, Germany in 1935 to escape Nazi persecution. I knew my father’s father only slightly as he lived in Milwaukee, WI, visited only occasionally, and died when I was only four years old. My grandmother Kaethe, on the other hand, lived to be 102, residing near my parents’ house for many of the last 40 years of her life. I had innumerable interesting conversations with her, beginning at a young age and continuing into my 40s. She was, on the one hand, kind to all her grandchildren, generally patient, and attentive to us. But occasionally she would lose her temper and shout at us with a strong German accent, which could be quite intimidating. She had an extraordinary vocabulary in English, was highly informed of current events, especially national and local politics, and spent much of her time reading books and political magazines. She was a lover of classical music and a competent pianist, who in her 60s and 70s, gave lessons to children in her neighborhood. She remained intellectually sharp, trading on the stock market and even learning to program a computer in her late 80s and early 90s. She had high standards, was quite strict about punctuality (once excoriating my sister when she showed up to an appointment at 3:17 pm rather than the designated 3:15 pm), and was hard to impress. She was not one to “ooh” and “ahh” over a child’s drawing or any other minor accomplishment. On the other hand, she did encourage all of us, often through unstinting praise of great achievements in the world at large. It may well have been from her that I first heard the phrase “Nobel Prize” when I was still a small child, and on rare occasions, when I talked to her about science, I recall her saying “maybe someday you will win the Nobel Prize.”
My grandmother was never comfortable as a Jew in German society at any time she lived there, although she was the physician to children of prominent gentile families in Berlin. Among her patients was the son of Magda Goebbels (from her marriage to Günther Quandt), about whom she spoke to me a few times. She described Magda as an “elegant, beautiful woman” who was quite pleasant prior to her marriage to Josef Goebbels, whereafter she visited no more. My grandmother did not foresee the Holocaust, even two years after Hitler’s rise to power, but did foresee that her three children (my father, Ernst, his older brother Frederick, and his younger sister Ruth) would be denied an education and would have no future in Germany. She was the prime mover in the family’s decision to leave. For a time, it was undecided whether they should emigrate to Palestine or to the United States. My grandfather actually visited Palestine, but found opportunities for physicians to be limited, and decided in favor of moving to America. This was reportedly contrary to my grandmother’s wishes. “He asked me where I thought we should go. I said ‘Palestine,’ so of course we moved to America,” she sometimes recounted. In the years that followed, she regretted that she could not take part in building the modern state of Israel. At the same time, she was relieved that her children did not need to fight in the wars that beset Israel from its founding.
Both of my parents were born in 1928, my father in Berlin, Germany, and my mother in Chicago, IL, USA. My father was said to have been a somewhat difficult child, often in conflict with his mother. However, he performed extraordinarily well at school, and at the age of 15 was enrolled in a special program developed by Robert Maynard Hutchins, then President of the University of Chicago, which permitted him to finish high school and college within two years and then go on to medical school. This he did, graduating as the valedictorian of his class, with a doctorate in medicine at the age of 21. He specialized in hematology, and in time became the pre-eminent academic hematologist of the 20th century. His scientific contributions extended to many areas of biomedical science, and he received numerous awards and distinctions in recognition of his work, including the Gairdner Prize in 1975, and election to the National Academy of Sciences and the Institute of Medicine in 1976. His most celebrated achievement was the 1962 discovery of random X chromosome inactivation in humans, made independently of Mary Lyon, who discovered it in mice. He made many other advances as well, in diverse fields of study, and he was to influence my own career dramatically.
My parents met at the University of Chicago, when my father was in his junior year of medical school. My mother was an undergraduate there, majoring in mathematics. They married June 15, 1950, and remained together until my father’s death in October of 2008. In addition to me, they had three other children: Steven in 1952; Earl in 1954; and Deborah in 1962 (Figure 1). Like me, Steve and Debbie became physicians; Earl became a successful businessman. Our early years were mostly spent in southern California, because my father moved there in 1959 to accept a position as Chairman of the Department of Medicine at the City of Hope Medical Center in Duarte. My mother worked as a homemaker for about 18 years, spanning most of my childhood. She then began a late career as a technical writer, retiring in her mid-60s.
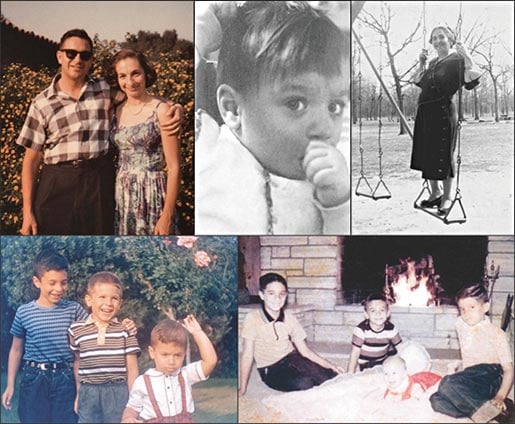 Figure 1. My early life and family. Clockwise from top left: My parents, standing in the front yard of our house in Arcadia, early 1960s (backdrop: a lantana bush often covered with skipper butterflies). Photograph taken of me at about 2 years of age, either before or after departure from Chicago for California. My maternal grandmother, Mary Fleisher (“Bubbie” to my sibs and me). My brothers and infant sister posing in front of the living room fireplace in 1962. From left: Steve, Bruce, Debbie, and Earl. My brothers and me, in the front yard of our home, ca. 1960. From left: Steve, Earl, and Bruce.
Figure 1. My early life and family. Clockwise from top left: My parents, standing in the front yard of our house in Arcadia, early 1960s (backdrop: a lantana bush often covered with skipper butterflies). Photograph taken of me at about 2 years of age, either before or after departure from Chicago for California. My maternal grandmother, Mary Fleisher (“Bubbie” to my sibs and me). My brothers and infant sister posing in front of the living room fireplace in 1962. From left: Steve, Bruce, Debbie, and Earl. My brothers and me, in the front yard of our home, ca. 1960. From left: Steve, Earl, and Bruce.
CA childhood and friends
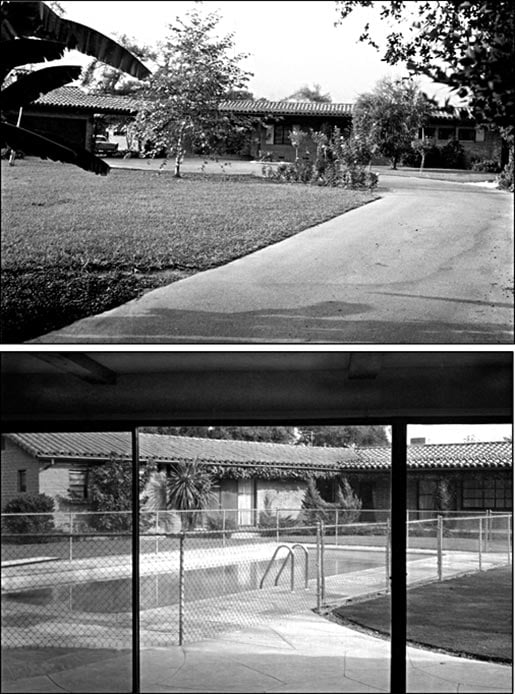
I was born in Chicago, IL, on December 29, 1957. But I have only fleeting memories of my first years there. My earliest memory is of newly fallen snow in the autumn of 1959. Seeing that the world outside had changed overnight, I asked “Who did that?” I also remember crying when movers came to take everything out of my room, as the family was relocating to California, where, from the age of two, I grew up in Arcadia, a northeastern suburb of Los Angeles. I have warm recollections of my childhood in California, and often have dreams in the setting of the adobe brick house where I grew up, with its orange terra cotta tile roof, one-acre yard, orchard crammed with fruit trees, guest house (which served as my room from the ages of 12–16), large front lawn (which we called “the field,”) and swimming pool (Figure 2). Generally speaking, I recall sunshine year round; heavy smog and hot weather during summer months; snow on the surrounding peaks and frost on the lawn in the mornings during winter; family vacations to national parks; and the fact that I had a strong interest in nature (particularly in animals). Over the years, I raised many animals (dogs, ducks, rabbits, mice, chameleons, tropical fish, turtles, and zebra finches), watching them and generally marveling at their behavior. As I grew up, I did lots of hiking (chiefly in the San Gabriel Mountains), birding, and bicycling. There were stresses to be sure: quarrels with my sibs and worries about deadlines in school, but nothing serious. On the whole I was very happy, even if I didn’t always realize it.

I began first grade early (at age 5), attending public school until I was 13 years old. I was then admitted to Polytechnic School, a college preparatory school in Pasadena, CA. The content of the curriculum at Polytechnic, and the pace at which we were taught, was quite different from what I had experienced in public school. It was something of a renaissance for me, and I regretted having wasted so much time beforehand. Not only was I taught more, and taught better at Polytechnic than at public school, but I also had exceptionally smart friends, and have remained close to some of them to this day. David Brittan, Paul Spiegel, Bob Kleinberg, John Taylor, and David Horowitz were among those friends, and with them, I could discuss almost any topic of the day, as they had a wide range of interests, spanning music, politics, literature, and science.
During three years at Polytechnic, I learned a lot about many things, and certainly changed a great deal, with some mental transformations literally occurring from one day to the next. For example, when I was 15 years old, I attended a performance of Bach’s St. Matthew Passion. My father had intended to go, but could not, as he needed to rescue my brother Earl, who had been bicycling from San Francisco to Arcadia, and had become exhausted on the way; hence I went in his place. I had been exposed to classical music in my home as a child, but it didn’t make any special impression on me. Yet I was completely electrified by this live performance of Bach’s great choral masterpiece, the first I had heard, and at once began listening to other choral music and collecting it on vinyl records, especially pieces by Bach, Handel, Vivaldi, Mozart, and Haydn. I remember arranging my curriculum in such a way that I could bicycle to school early in the morning (a distance of eight miles), complete my classes by late morning, and then bicycle home to listen to a radio concert of Renaissance music in my father’s study. Later I would do my homework (generally until quite late at night, and usually accompanied by Bach). I have remained a music lover ever since, ultimately settling on Bach as the center of my musical world. But although music became a deep and enduring interest of mine, I never learned to play any musical instrument proficiently.
Early interest in science
From childhood, living things appealed to me aesthetically. I could relate to them, and I was amazed by their similarity to humans. I was aware that life forms had changed continuously on earth over many millions of years, and concepts of genetic variation, natural selection, and inheritance were second nature to me, even from the time I was in elementary school.
In hiking and birding, and in looking at microbes through a microscope (one of my favorite diversions at some point), I enjoyed the observational side of science, which is where scientific inquiry normally begins. It was during high school, in the rich intellectual environment I have briefly described, that I first began to wonder about unanswered questions in science. The ability of inanimate molecules to assemble themselves into living matter was something that I found immeasurably interesting. My oldest brother, Steve, once introduced me to a quotation he attributed to Camus: “Life is the disease of matter.” Probably it was actually from Goethe, who as I later learned, wrote “Viewed from the summit of reason, all life looks like a malignant disease.” The idea resonated with me, not in a morbid sense, but because life was a process that compelled matter to do its bidding, co-opting it more or less automatically, and endowing it with special properties not seen in the inanimate world. In the early 1970s, I read the second edition of James Watson‘s “Molecular Biology of the Gene” (also a gift from my brother Steve, then in college at the University of California at San Diego) from cover to cover, and understood for the first time how DNA specifies the synthesis of proteins with specific functions, which in turn permit DNA replication, the development of a complex organism, meiosis, and the propagation of species. I became even more eager to participate in the study of living things, and to be an experimentalist rather than merely an admiring witness.
In high school and in college, I began to work in my father’s laboratory, and it was at this stage that I began to do authentic research. I was guided by my father and his interests, and learned to assay erythrocyte enzymes, and to characterize their electrophoretic mobility. One of the enzymes I studied most was glutathione peroxidase, unusual because it was a seleno-enzyme. At one point, striking out on my own a bit, I decided to see whether bacteria might have glutathione peroxidase, and whether it might be subject to biosynthetic control based on the availability of selenium, like the genes of operons, which had been described not too long before by Jacob and Monod. Lysing E. coli, I first found that there was no glutathione peroxidase activity. My father suggested I should add sodium selenite to the culture. I did so, and found considerable activity, which I interpreted as a success: I believed the selenite had induced synthesis of the enzyme. “Boil it,” said my father. The activity remained and was even slightly increased. Moreover, inorganic sodium selenite (and even more so, seleno-cysteine) exhibited catalytic activity. When seleno-cysteine was incorporated into the protein, this activity was enhanced many thousand fold. We proposed a reaction mechanism based on the catalytic properties of selenium in distinct oxidation states. We also found a polymorphism of erythrocyte glutathione peroxidase activity in members of my own family, and an electrophoretic polymorphism in others. I began to think as a biochemist, and to some extent, also as a geneticist, but only in a rather elementary way at that stage.
My father as a role model
The foregoing vignette tells something of how my father influenced me. I was thrilled when he received the Gairdner Award, and when he was elected to the National Academy of Sciences, both of which occurred during my teenage years. Indeed, I sought to emulate him and took his advice seriously. When he suggested I read “Arrowsmith” by Sinclair Lewis, and “The Microbe Hunters” by Paul de Kruif, I did so, and just as they had earlier affected him, they also affected me. Among the most important pieces of advice he gave me was to go to medical school. The explanation he gave was that disease often reveals new and important biological principles; also, as a physician in training, one acquires broad knowledge of anatomy, physiology, histology, pathology, and pharmacology: sciences that come into play in understanding many biological phenomena. Throughout his life, my father was an advocate of working on “something important,” rather than what he saw as esoteric topics. Generally speaking, he preferred authentic clinical research (with patients) to research with mice, and research with mice to research with flies or other distant model organisms. Notwithstanding his strong genetic and evolutionary orientation (which we shared), he tended more toward applied research than I myself did. And his judgment as to what was important rested chiefly on his medical experience, which was quite extensive.
With a strong focus and a sense of mission, I completed high school early (at the age of 16) and enrolled at the University of California at San Diego (UCSD), in Revelle College. I had taken certain examinations that gave me “advanced placement” credit, and I pursued an ambitious schedule in college. I worked through two summers, and graduated at the age of 18. I was a good student, but not an outstanding one. As a teenager and even for some years beyond, I was a bit emotionally volatile, often distracted by love interests, and generally speaking, in too much of a hurry. I took on an enormous course load, wishing to get through college quickly, go on to medical school, and become a scientist. It was an exciting time in biology, after all, with extraordinary advances in molecular cloning beginning to dawn. I was aware of what was going on, and impatient to be part of it. To my friend high school friend David Brittan (also my roommate for a time in college), I remember saying with frustration “The train of science is leaving without me.”
But despite the hurry, I did learn a lot, and only realized how much later on. I had a truly stellar introduction to genetics in the laboratory of Dan Lindsley (Figure 3a), a distinguished Drosophila geneticist interested in spermatogenesis and spermiogenesis in the fly, among other topics. In his lab, I attempted to map the gene for hexosaminidase (a project inspired by my father, who had studied this enzyme in humans, where hexosaminidase deficiency causes Tay-Sachs disease). When I was 18, I spent the summer working in the lab of Abraham Braude, most immediately with his postdoctoral fellow Arthur Friedlander (who later became well known for his studies of anthrax lethal factor). In Braude’s lab, I first heard the word “endotoxin,” also known as lipopolysaccharide, or LPS. I understood it was pyrogenic, capable of activating leukocytes, and also heat stable (which made it a particular problem in our studies of chemotaxis). All glassware had to be heated to 180°C to destroy contaminating LPS. I also knew that Braude had tried to passively immunize humans against LPS to protect them against Gram-negative septic shock. But my interest in LPS was casual and tangential at that stage, probably because its biomedical importance was still an abstraction to me. I had never seen a patient with sepsis, and didn’t grasp what a serious clinical problem it might be. The question of an LPS receptor did not occur to me at that time. I did not remotely imagine that searching for it would form the core of my Nobel Prize-winning work two decades later.

After completing college, I had still more exposure to genetics in the lab of Susumu Ohno (Figure 3b), a friend and colleague of my father at City of Hope Medical Center. I worked with Ohno for about nine months, and spent still another summer with him during medical school. A famous mammalian geneticist, Ohno had demonstrated that the Barr body observed in cells of female mammals was a condensed X chromosome. He had also developed the thesis that evolution depends upon gene duplication. He had written extensively on the phylogeny and origin of sex chromosomes. And he had observed that the genetic content of the X chromosome tends to be strongly conserved in all mammalian species. In short, he was a remarkable theoretical biologist at a time when experimental tools were not nearly what they are today.
Ohno inclined toward immunology after spending a sabbatical at the Basel Institute for Immunology. He hypothesized that major histocompatibility antigens act to anchor organogenesis-directing proteins. He adduced considerable experimental evidence supporting this hypothesis … which we know today was entirely incorrect! The time I spent in Ohno’s lab, working on this very subject, was quite enlightening for me from many points of view. In hindsight, perhaps the most important – if brutal – lesson was that even extremely intelligent scientists can deceive themselves if they embrace hypotheses too passionately. But I also learned much about immunology as it was understood in the mid-1970s.
I applied to several medical schools, confident that I would be admitted to most of them. I had good grades, extensive experience in laboratory research, several publications (including one first-author paper in Cell), and MCAT scores in the top percentile. But in the end, I was admitted to only one medical school: the University of Chicago. All others declined my application, perhaps because I was so young, and perhaps because I was unreservedly interested in science rather than clinical medicine. So I must be grateful to the University of Chicago for the chance it gave me. Luckily, it was (and remains) an outstanding institution. In the fall of 1977, I moved to Chicago and began my studies there at the age of 19, the youngest member of a class with more than 100 students.
Medical school
My first impression of Chicago was that it was a “real” city compared to the southern California suburbs I had known. But it was also somewhat dangerous (where I lived, at least), and comparatively unwelcoming. The Chicago winter was something for which I was not prepared. I arrived dressed as a Californian, and nearly froze when I tried to walk the mile from the University to my apartment during the first winter storm, in high winds and sub-zero temperatures. As I remember, my ears were insensate for about an hour, and I initially feared they would become gangrenous and fall off! I recall the first years as quite challenging. Anatomy and neuroanatomy were especially tough, requiring spatial memorization of a type that was unfamiliar to me, but in the end, both classes were rewarding. My classmates were for the most part smart and competitive, and I had to struggle to do well. I particularly enjoyed histology, physiology, and microbiology classes.
When it came to my introduction to clinical medicine, the work was more demanding than any I had ever known. Ultimately this was beneficial, because I came to expect equivalent discipline of myself and others when I worked in the laboratory. But I did not get the same joy out of clinical work that I did from laboratory work. I was often uncertain as to whether particular therapies and practices had a sound rational basis, and when patients got better, I did not always feel I could claim credit for it. After all, there was no control group. And very rightly, there was no leeway for experimentation on the wards. I did manage to work for some months in the laboratories of Patricia Spear, an outstanding young herpes virologist, and Barry Arnason, a neurologist with a strong interest in multiple sclerosis. But overall, there was little time for research, and I began to miss the lab.
In 1980, when I was 22, I married Barbara (Barbie) Lanzl, then a dental student at nearby University of Illinois. She was three years older than I, and had previously been married to my friend and medical school classmate Jiri Sonek. The marriage lasted 8 years, with phases in Chicago, Dallas, New York, and Dallas again, before ending in divorce. We had three bright, healthy sons, two born in Dallas and one in New York.
Internship and residency
I graduated from the University of Chicago in 1981, and again acting according to my father’s advice, decided to spend at least a year or two in residency, learning more about clinical medicine. I was matched with the internal medicine internship program and the neurology residency program at UT Southwestern Medical Center in Dallas. Dallas was my top choice, and arguably offered the finest clinical training in the country. It was known for giving interns and residents considerable responsibility and autonomy. During the year of my internship, for example, two interns and one resident would run the internal medicine emergency room alone through the night, with no attending physician present. And when necessary, medicine interns were expected to perform fairly invasive procedures, including intubation, placement of subclavian lines, cardiocentesis, insertion of chest tubes: in short, whatever was required, particularly during emergencies (which were not uncommon).
My father’s twofold rationale in suggesting an internship and residency rather than a laboratory fellowship had been that I would become “a finished doctor;” also that I would have something to fall back on in the event that research did not pan out for me.
As to the first point, I believe he was correct. At least the year I spent in internal medicine internship taught me many things I hadn’t known as a senior medical student. By the time I had finished, I feared no medical emergency; I felt secure in doing whatever needed to be done. But I also knew that medicine was not for me. I badly missed research and longed to return to it. I felt obligated to finish at least one year of neurology residency and did so. I learned quite a bit about neurology, which sometimes helps me to the present day in assessing mutations in mice. But perhaps the residency year gave me a bit more clinical training than I ever wanted.
As to the second point, my father was certainly being prudent. But to my recollection, I was supremely confident of success in science. I felt that no matter which lab I joined, and no matter what project I undertook, I would prevail, given the interconnectedness of biological processes, and my overall knowledge about how they worked. I was dead certain of my skills. I had a “feeling” for proteins and how to isolate them. I had a solid grounding in genetics and in immunology. And, true to my father’s earlier advice, medicine had taught me much about how an organism functions: far more than I would have learned had I gone to graduate school. Perhaps I was brash, but in recalling my mental state at the time, I would say I was imbued with the enthusiasm of youth, and felt invincible
Rockefeller University and an early success
I was 25 years old when, on the evening of July 4, 1983, Barbie and I arrived in New York City together with our infant son Danny, who had been born in Dallas. I remained in New York for three years, and during the second year, our second son Elliot was born. Most of the time, the four of us lived on the 11th floor of a high-rise apartment building across the street from the Rockefeller University campus, with a nice view of the East River.
I had joined the Cerami laboratory at Rockefeller. There I began to work on cachectin, an unidentified factor expressed by macrophages in response to activation by LPS, and defined by its ability to suppress lipoprotein lipase synthesis in adipocytes. The prevailing hypothesis in the laboratory at that time was that cachectin was responsible for wasting in chronic diseases such as tuberculosis, trypanosomiasis, and cancer. From the start, I was skeptical about the idea that a single factor could explain all cachexia, given the diversity of inciting causes. At least, there was no strong reason to think so. But I did see early on that a single factor responsible for suppression of lipoprotein lipase in fat cells was secreted by LPS-activated macrophages. Moreover, I saw that this factor was a protein, and I felt it could be purified.
At the time I started work in the lab, no progress at all had been made in isolating cachectin. Cachectin activity had been ascribed by my predecessors to a protein with a molecular weight of 70 kilodaltons (almost surely contaminating serum albumin). Moreover, they had purportedly excluded tumor necrosis factor (TNF ) as a candidate mediator by exchanging material with the laboratory of Lloyd Old, who had discovered TNF and was then still trying to isolate it. They had erred in reaching this conclusion.
Within a year, I purified cachectin to homogeneity, raised an antibody against it, and affinity purified the antibody. I hypothesized that whatever role cachectin might have in cachexia, it was likely involved in endotoxic shock, as it was produced in large amounts when macrophages were activated by LPS, constituting about 2% of their secretory product. Moreover, I found that purified cachectin was capable of killing mice when as little as 20 micrograms was administered by an intravenous route. One evening, I passively immunized 5 mice against cachectin, and gave pre-immune globulin to 5 control animals. I had formally randomized the mice to the two treatment groups, and I waited for several hours to give the antibody time to distribute through all extracellular fluid compartments. I then challenged the animals with a carefully chosen dose of LPS: a low LD100. To my delight, I found the next morning that passive immunization protected all 5 recipients against LPS-induced death, while all 5 control mice had succumbed! I repeated the experiment many times and with many permutations, using hundreds of mice, and found the result to be robust and reproducible. I was thrilled because I knew I had isolated one of the key endogenous factors mediating the lethal effect of LPS. The inflammatory properties of cachectin, rather than an ability to induce wasting, proved to be immensely important.
Several months passed before I could determine the N-terminal sequence of cachectin, as my initial purification strategy left the protein N-terminally blocked. I needed to repurify it using an entirely different procedure. Eventually, 17 residues were called by Edman degradation. Initially, I believed cachectin was a novel protein. Its sequence did not appear in the rudimentary protein databases that existed at that time. However, I was soon alerted by the Ulevitch group at Scripps that my purified cachectin, sent to them for separate purposes, had high tumor necrosis factor activity. I directly compared the sequence of human TNF, only recently isolated and cloned at Genentech, to the sequence of mouse cachectin. I saw strong homology. Cachectin was the mouse orthologue of human TNF.
This worked to my advantage, perhaps, in that there was already great interest in TNF as an anti-neoplastic protein. Yet I had then shown that the protein also had toxic, inflammatory properties. In time, TNF became one of the most studied proteins in biomedicine, and attempts to block its activity with antibodies (or with recombinant proteins such as my group later developed) bore more fruit than attempts to administer it.
During my three years at Rockefeller, I published many papers, including several in high-ranking journals, and made a substantial name for myself in the rapidly developing cytokine field. Soon, many thousands of other publications cited my work. I obtained my own funding and was promoted to the rank of Assistant Professor at Rockefeller in 1985, but this post did not carry true autonomy. I wanted a lab of my own. When I was invited by Joe Sambrook to join the Howard Hughes Medical Institute (HHMI) at UT Southwestern Medical Center, I eagerly left New York to forge my own future.
Dallas and the Lps locus
Returning to Dallas in 1986, one of my first and best decisions was to hire Betsy Layton as my secretary. She was 19 years old at the time, and had just moved to Dallas from her native Pennsylvania, attracted by the image of Texas and by opportunities for work. When I interviewed her, she struck me first of all as nice, cheerful, smart, and friendly; also as someone who had a strong will to work and to do a good job. I was right on all of these calls. Meeting Betsy was one of the luckiest things that ever happened to me, because she was truly devoted to me over the many years that followed. She helped me to set up my lab, offered advice with appropriate tact and candor, and became a true partner in my professional life. In time, she became an administrative assistant; then an administrative manager. She accompanied me from Dallas to Scripps, and then back to Dallas, where she still works with me today. I hope this will always be the case. In the end, choosing outstanding people and guiding their work well becomes far more important than the work a biologist does with his own hands.
When I first settled in Dallas, I began to pursue several topics related to TNF. The most practical of these was to develop a means of blocking TNF activity in vivo. I engaged with this challenge partly because I knew that it would be useful to do so both in chronic inflammation and in septic shock, and partly because I wished, in the “pre-knockout” era, to see what a chronically TNF-deficient animal would be like. David Crawford, an MD/PhD student, and Karsten Peppel, a postdoc in my lab, developed a chimeric molecule in which the ectodomain of one of the TNF receptors was fused to part of an IgG heavy chain, including the hinge and Fc region. As we had hoped, this yielded a non-antigenic, stable, and extremely potent neutralizing reagent that could be administered to mice (or humans) to block TNF activity in vivo for long periods of time. We patented this protein, and eventually sold the patent to Immunex, which in turn was acquired by Amgen. Today, the molecule we invented is marketed as the drug Enbrel, and is used as an effective treatment for rheumatoid arthritis and several other inflammatory diseases.
On a more basic level, I focused my laboratory on the question of how TNF biosynthesis might be regulated. I viewed this as tantamount to the question of how the inflammatory response might be regulated as a whole, in that it was already apparent to me that TNF was an excellent marker of the inflammatory response. I had earlier shown, for example, that anti-inflammatory drugs such as glucocorticosteroids could entirely abolish TNF biosynthesis in response to LPS, and felt that this explained a large part of their inflammatory effect. I deduced that TNF biosynthesis was regulated both at transcriptional and translational levels. TNF gene transcription was dependent upon NF-ƙB, a factor soon seen to induce many cytokine genes. Translational repression depended upon a sequence my colleague Daniel Caput and I had discovered in the 3′-untranslated region of the TNF mRNA, consisting of overlapping and interleaved octamers with the sequence UUAUUUAU. A similar sequence, I had noted, was found in many cytokine encoding mRNAs. Activation of the macrophage by LPS overcame translational repression. Both Véronique Kruys and Jiahuai Han in my lab closely studied these phenomena, but we did not succeed in elucidating the biochemical details of translational regulation during those years.
The more central question as to how macrophages became activated by LPS remained elusive. At the core of the question was the issue of the LPS receptor. TNF was made in large amounts in response to LPS. Hence, I was inquiring into LPS signaling when I attempted to measure responses of the TNF gene and mRNA. Yet there was no understanding as to how LPS was perceived by cells in the first place. And long before most others in the innate immunity field, I regarded the question as one of the most fundamental and important in all of immunology.
After all, since microbes had been identified as the causative agents of infectious diseases in the mid-1800s by Pasteur and Koch, nobody had determined which receptors recognize their presence during infection. This was the first level of self/non-self discrimination by the immune system. LPS mimicked infection in all its complexity quite closely, and potentially, the LPS receptor might be responsible for all events that transpired during a real infection. In finding the LPS receptor, we could hope to know at least one key sensor used by the innate immune system to recognize microbes. Perhaps it might be a member of a receptor family; perhaps we would gain insight into how all microbes were detected within the first minutes following inoculation.
I was aware of two strains of mice that were specifically refractory to LPS, because of mutations affecting the so-called Lps locus. These were the C3H/ HeJ and C57BL/10ScCr strains. For each, a closely related strain (C3H/ HeN and C57BL/10ScSn, respectively) served as a control with normal LPS responses. And mice of both strains were known to be susceptible to Gramnegative infections. In my mind, these mutations abrogating LPS responses grew constantly in stature. What protein did they affect? Was it indeed the LPS receptor?
Positionally cloning the LPS locus was beyond my capabilities during the 1980s. I had isolated TNF in a lab in which molecular cloning had never been practiced, and I had to teach myself almost everything in the way of basic molecular biology methods. I understood what needed to be done, but did not at that time feel secure in tackling such an immense problem. I tried instead to look for a difference between control mice and mutants at the protein level. And I attempted to raise an antibody against the proteins of a WT mouse in a resistant animal (or vice versa) to discover a “missing” or altered protein in mutants. I also tried to use expression cDNA cloning to find a gene product that could rescue the LPS-resistant phenotype. We tried, too, to use insertional mutagenesis (with a retrovirus) to inactivate the LPS locus in heterozygous mice made by crossing C3H/HeN animals to C3H/ HeJ animals. But as all of these approaches to finding the LPS receptor were unsuccessful, I began to think positional cloning might be the only way to go.
I began to pursue this strategy actively in 1993, when Christophe Van Huffel came to my laboratory from Belgium to work as a postdoctoral associate. He had a background in yeast genetics, which made me think he would be a good person to isolate the YA C clones we would need to build a contig across the critical region. We began mapping the mutation in C3H/ HeJ mice to high resolution, ultimately including a total of 2093 mice in our meiotic analyses. We became stronger in our molecular skills, and gradually, more people were incorporated into the effort, until the entire lab had a single focus. Among these were Alexander Poltorak and Irina Smirnova, who worked devotedly and tirelessly to find the mutation, year after year. They were exceptionally able colleagues. Betsy too joined the effort, and helped to read sequence, organize data, pick colonies for sequencing. Alexander, Irina, and Betsy were as zealous as I was; they simply refused to give up.
I have not directly mentioned it to this point, but I tend to be an obsessive person in many ways. I am even somewhat proud of this, although in some circumstances it has brought me considerable suffering without tangible rewards. Finding the mutations that abolished LPS responses became one of the most gripping obsessions of my life. For years, it occupied much of my waking consciousness and sometimes my dreams as well. Once, in 1997, while staying in a hotel in the San Bernardino Mountains, I awoke from a dream in which I was sure I had come to understand what gene was affected by the mutation. I hastily wrote the name of the affected protein on a scrap of paper and happily returned to sleep. In the morning, I saw that the protein was complete nonsense; it did not exist (though from the suffix “ase,” I could tell it must have been an enzyme)!
Positional cloning of the LPS locus was the toughest challenge I had ever faced for three reasons. First, the critical region was gigantic: by far the largest ever tackled in the mouse. 24 BAC clones and one YA C clone were needed to span it, and even then there was a small gap we never closed. We know today that the genomic interval was at least 5.6 million base pairs in size. To have such a region within which crossover was apparently forbidden was simply bad luck, and at that, we needed to explore about 90% of it before we found the mutation. Second, we had limited sequencing power. We began sequencing with radionuclides (mostly 35S), loading gels by hand, drying them, and reading sequence ladders on X-ray film. Later we used slab gels with semi-automated fluorescence-based reading of sequence. But we never had capillary sequencers at our disposal. This made it difficult for us to examine the region for genes. Third, the methods used at the time to find genes were primitive. We began with exon trapping and hybridization selection. Only by the mid-1990s were complex expressed sequence tag (EST) databases available. Matches between genomic DNA in our interval of interest were sought in the EST databases using an algorithm called BLAST. And BLAST searching became a major chore, because we had insufficient local computing power. I taught myself to program in Perl in order to manage the many thousands of sequence files, and the output BLAST data in a semi-automated way. For years, no meaningful hits emerged. There were only hits derived from pseudogenes, which nonetheless had to be cloned, sequenced in full, and run to ground, so that we might be sure we could find no mutation distinguishing C3H/HeJ from the control strain C3H/HeN.
Pressures, criticism, and anxiety
Both personal and professional pressures weighed heavily on me during the years of the LPS cloning project. My youngest son, Jonathan, was born in 1987, and Barbara and I separated in 1988. The divorce proceedings were contentious, and went on for years, involving innumerable depositions, preliminary hearings, and a jury trial, ultimately leading to a joint custody order. Of course, life did not immediately normalize thereafter. Particularly during their teenage years, my sons were each difficult to manage. I am happy to say that all three are extremely close to me today. It was a special joy to bring Danny and Elliot to Stockholm for the Nobel Prize ceremony (unfortunately, Jonathan was unable to attend). But in the mid-1990s, there was a lot of stress, and tough times at home coincided with the toughest phase of the cloning work.
While the importance of finding the LPS mutations was obvious to me, it was less obvious to other people. Years elapsed with no publications to show, and some of my colleagues thought that we did not entirely know what we were doing; some believed that the gene might, in the end, not be particularly illuminating; all believed that we were taking a terrible risk. Among these was my father, who urged me to diversify my portfolio and not to “put all your eggs in one basket.” I ignored his advice on this occasion; I simply couldn’t abandon or in any way diminish the intense focus on the LPS locus, because I knew we wouldn’t find the mutations if I did. In hindsight, I am sure I was correct about this.
At least two other groups were attempting to find the same gene. One was the group of Danielle Malo, in Montreal; another was the group of David Schwartz in Iowa. I worried in particular about Malo’s effort, because she and her close colleagues had an impressive record in positional cloning. At one point, I offered to collaborate with her, but the offer was declined, which meant we were formally in a state of competition.
HHMI was increasingly impatient with me. Over the 14 years I worked as an HHMI investigator, my program was reviewed by the Medical Advisory Board five times. After our final review, in April of 1998, I was told I would be funded through August of 2000, but not beyond. This was disappointing, since we had indeed made progress, and I felt that success was imminent. But their indulgence was at an end.
I felt we must continue come what might, and we did so, though some of the postdoctoral fellows in my group, Christophe Van Huffel, Mu-Ya Liu, and Xiaolong He, left to find other laboratories, as did some of the technicians, who also sensed impending defeat. Hence, it was an anxious time. Most of the critical region had been explored in depth. Surely we would find the gene soon. Or else, perhaps, we had made a mapping error and were looking at the wrong part of the chromosome. Were we on the cusp of success? Or had I been a fool to reject wise counsel, and would our efforts end with nothing to show for them?
Success
The gene we had struggled with for five years was discovered very suddenly one night: on the 5th of September, 1998. I was reviewing the day’s BLAST results as they returned from NCBI and from our own server where we had begun to run BLAST locally. I was shocked by what I saw, as there had been a long dry spell, and here was one … then two … matches with a real gene: Tlr4. Almost certainly it was not a pseudogene based on the quality of the hit. That in itself encouraged me to think that this was the “holy grail” we had been seeking: here was an authentic gene, and only a small amount of genetic material that remained unexplored. Moreover, a good story could immediately be made about the candidate. It appeared to be a cell surface receptor with a single membrane spanning domain. The ectodomain was rich in leucine, which might be expected to bind hydrophobic molecules like the acyl chains of LPS, and was similar in overall structure to CD14, earlier shown to be required for sensing LPS molecules. It was a member of a family of proteins related to Toll, which in Drosophila was known to have both an immune and a developmental function. And on the cytoplasmic side, it was similar to the IL-1 receptor, well known to deliver an inflammatory signal.
I related these facts breathlessly to my father (who was somewhat nonplussed, I must say), and to Alexander and Ira, who were very much more excited, sensing as I did that our battle might be nearing an end. But we would not be on firm ground until we were able to find a mutation distinguishing the C3H/HeJ strain Tlr4 from the C3H/HeN strain Tlr4, and the C57BL/10ScCr strain Tlr4 from the C57BL/10ScSn strain Tlr4.
The very next day, we attempted to amplify the cDNA in all these strains by long-range PCR. Alexander succeeded in so doing using cDNA from the C3H/HeJ, C3H/HeN, and C57BL/10ScSn strains, but failed to do so using cDNA from the C57BL/10ScCr strain. We therefore concentrated first on the C3H/HeJ and C3H/HeN sequences, which Alexander established by shotgun sequencing. Within days, we saw the mutation for the first time: it was a single base pair change predicting the substitution of a conserved proline for a histidine in the cytoplasmic domain of the molecule. We later established that the gene was deleted in the C57BL/10ScCr strain. And this eliminated any remaining doubt that TLR4 was essential for LPS signaling.
I called James Gavin, my HHMI liaison, to tell him what we had done. In reply, he made no comment on our findings, but emphasized that the decision of HHMI was irrevocable. For my own part, I was so elated that we had made a truly great discovery that the blow was much softened. We submitted our work to Science, where it was promptly accepted for publication, and meanwhile, presented it at four international meetings, where it was received with acclaim (Figure 4). By now it has been cited over 4,000 times: more than any other paper in the innate immunity field.
Most exciting of all, perhaps, was the fact that the LPS receptor was indeed part of a protein family, and it could easily be imagined that other TLRs detected other molecules of microbial origin. In time, this proved to be the case. And in due course, we were able to show that TLR4 did in fact engage LPS in order to detect it. That is, TLR4 did not act as a signaling intermediate as its Drosophila homologue did, but as the receptor per se. We had opened a window into how mammals perceive microbes, and a great many laboratories began to study the question of how TLRs signal to initiate the inflammatory response, and how they might be involved in the pathogenesis of sterile inflammatory diseases.
La Jolla (2000–2011) and ENU mutagenesis
In 2000, I relocated my laboratory from UT Southwestern to The Scripps Research Institute (TSRI). Eager to dissect innate immunity in mammals, I invested heavily in ENU mutagenesis as a means of creating new and interesting phenotypes in the mouse. The plan was to create many exceptions to the norm using a random process, thus acquiring fresh paradigms of importance equal to the C3H/HeJ and C57BL/10ScCr mice (which bore spontaneous mutations, and taught us so much about microbe sensing). In this way, I felt, we could take innate immunity apart gene by gene, finding all the parts of an enormous puzzle, and do so without hypotheses and their attendant biases.
I chose the classical genetic route because I knew the mouse genome would soon be sequenced and annotated; hence positional cloning would become much easier. All candidates would be known within all critical regions, and only mapping would be required to exclude most genes from consideration. Never again would it be necessary to build a contig and explore it for genes: the two toughest parts of positional cloning. Moreover, sequencing candidate genes was becoming progressively easier. Capillary sequencing was replacing the slab gels, and there was great potential for automation. In time, I foresaw, entire genomes might be sequenced all at once. The main burden would be to generate phenotype for study: in our case, immunological phenotype.
But until sequencing technology had advanced further, the crucial point was to cover critical regions quickly and efficiently. Toward this goal, Yu Xia, an exceptionally skillful programmer in my group, wrote software that would analyze genes within a specific genomic interval, mask the sequence to hide common repeats, and design optimized primers for amplification and sequencing. He also wrote code that would permit mutation identification: a human observer no longer needed to search through trace files to find them. This was a system that no other laboratory possessed, and using it, we were able to positionally clone as many as 50 mutations per year (as compared to one mutation in five years only a decade earlier).
As “next generation” sequencing became a reality, we modified Yu’s system to validate mutations identified within the whole mouse genome. At last, only the most cursory mapping was required to define the location of a mutation, whereon it could then be found by machines that sequenced DNA a million times faster than we were able to sequence it in former times. This brings us to the present state of the art. My colleagues and I have found hundreds of mutations responsible for phenotype, many of them affecting immune function. Like all mechanists and reductionists, we regard the innate immune system as a highly complex machine. Surely we have far to go to understand its workings as we might understand those of a pocket watch. But just as surely, the innate immune system is a machine, and one day it will universally be seen as such.
Over the 13 years that elapsed between our discovery of the LPS receptor and the announcement of the Nobel Prize, I made a number of close friends in the innate immunity field. Shizuo Akira made enormous contributions, as he and his colleagues used reverse genetics to dissect the signaling pathways incorporating the receptors, adaptor proteins, kinases, and transcription factors that lead to activation of the inflammatory response. Jules Hoffmann, who with his colleagues had discovered the immunological role of Toll in the fruit fly, went forward to analyze the Drosophila imd pathway (evocative of the mammalian TNF signaling pathway, and used by the fly to detect Gramnegative bacteria) and the upstream proteins that activate Toll and imd signaling. Jules’ group, Shizuo’s group, and my own began to study antiviral defense in insects and mammals, collaborating in an open and congenial manner. I developed the highest respect for their scientific acumen and integrity. Jules and I, in particular, spent pleasant days together over the past decade hiking, discussing politics, and talking about history, science, and people.
I also developed strong friendships at Scripps, none stronger than with Argyrios (Ari) Theofilopoulos. Ari had worked for many years to understand autoimmunity, especially as represented in systemic lupus erythematosus (SLE). A part of the premise for searching for the LPS receptor was the likelihood that it might ultimately explain sterile inflammation. In fact, TLR7 (and conceivably other TLRs as well) turned out to have an essential facilitating role in SLE, at least as modeled in the mouse. Ari and I had numerous discussions about this, and collaborated quite extensively on studies of autoimmunity using TLR mutants and other mutants generated using ENU mutagenesis.
In time, our work in the innate immunity field was recognized by some major prizes: first the Robert Koch Prize (2004), then the Gran Prix Charles Leopold Mayer (2006), the William B. Coley Award (2006), the Balzan Prize (2007), the Albany Prize (2009), the Will Rogers Institute Prize (2009), the Shaw Prize (2011), and others in between. In 2008, I was elected both to the National Academy of Sciences and to the Institute of Medicine. Each of these distinctions was an occasion to celebrate, often with family, friends and colleagues in attendance (Figure 5). I was happy that my father was alive to mark most of these milestones with me. But to the sorrow of all my family, he passed away in October of 2008, after a year-long battle with mantle cell lymphoma.

The present, the future, and advice to others who may follow me
By March of 2011, I had decided to return to Dallas, and I had a dual appointment at Scripps and at UT Southwestern on October 3 of that year. By the time the Prize was awarded in December, I had completed my relocation. I have begun to build a Center for the Genetics of Host Defense, in which faculty with a forward genetic orientation, working with diverse model organisms, can join together to study the protective mechanisms that have developed to combat infection. They may also study autoimmune and autoinflammatory diseases: the destructive legacies of immunity, which was itself imposed on us by the enormous selective pressure that infections represent.
I must admit that many questions in biology appeal to me, and at times, it is difficult to maintain an intense focus on the immune system, important though immunity is. The great epiphany for me in solving a single, critical question – the conundrum of LPS sensing – was the embrace of genetics. This is a golden age of genetics, in which one may find the cause of monogenic phenotype almost immediately. The destruction of all genes in the mouse by chemical and/or insertional mutagenesis and gene targeting will soon be a reality. At this writing, the ability to clone mice from somatic cells, and the availability of haploid embryonic stem cells for genetic experimentation, also promise to accelerate progress in biology. When I first began working in my father’s laboratory, none of these technologies were foreseen by anyone then alive. It is fair to say that mankind’s understanding of living things has advanced more during the past 50 years than it did during all of history before. I have been a direct witness to most of this progress. So have many others who are older than I, and have been practicing scientists for a longer time. This is to say that we live in an exciting era, and should feel lucky about it.
The Nobel Prize gives an opportunity for introspection, and to conclude this brief autobiography, I offer some thoughts that arose in my mind during the last few months, mostly prompted by specific questions from students, journalists, family and colleagues. “To what do you attribute your success?” they want to know. “How does one win a Nobel Prize?”
In my case, the path to the Nobel Prize began with a love of nature and an earnest curiosity about the phenomenon of life. I was lucky to have an understanding father, who was himself a distinguished scientist. He gave me excellent advice during my formative years. But this might have helped little had I not been industrious, enthusiastic, and optimistic in pursuing a career in biology. I did not worry or make contingency plans to be used in the event of failure; I ran blithely ahead with confidence that success would come to me.
I chose an undervalued question, viewing the mystery of the LPS response as the key to understanding inflammation and the recognition of microbes – and I based this assumption on a strong and well-studied phenotype. While the LPS receptor did not turn out to explain everything about how inflammation works, it did give major insights. The choice of a question is a matter of scientific taste; a matter of recognizing what is interesting and important. Such taste is partly innate and partly learned. To the extent that I learned scientific taste, I did so by studying medicine.
Much later than I might have, I embraced genetics, preferring its unbiased character to hypotheses, which often lead scientists astray, mainly because they don’t like to be wrong. I do not eschew hypotheses. But not all scientists have the integrity to test them as they should be tested. Only the very best scientists do. The others try to “prove” their hypotheses. And this unfortunate fact is the cause of much error, wasted time, and wasted resources.
I have also learned that it is important to choose outstanding colleagues. Not only those who will frankly criticize one’s work or provide fresh insight, but also those with whom one works directly, as a mentor or supervisor. Recognizing talent, acquiring it, nurturing it, and retaining it is among the hardest things a scientist must do. It is a skill quite separate from scientific ability per se. It will usually determine whether the scientist succeeds, because none of us can accomplish much in isolation.
Finally, and maybe most important of all, my principal achievements resulted from addressing a single problem with relentless obsession. Once convinced that something could be achieved, I was not deterred by discouraging advice from others. I would suggest the same course to anyone. In the event of failure, one may always select another problem and begin anew. That is what I would have done had I failed. As it happened, it was unnecessary.
Nobel Prizes and laureates
Six prizes were awarded for achievements that have conferred the greatest benefit to humankind. The 14 laureates' work and discoveries range from quantum tunnelling to promoting democratic rights.
See them all presented here.
